Unlock Limitless Possibilities with HALO AI
HALO AI has you covered. Whether you’re looking to solve image analysis challenges in segmentation, classification, or phenotyping, the advanced deep learning networks of HALO AI are easily tuned for your specific application with the train-by-example interface.
Diverse and Powerful AI
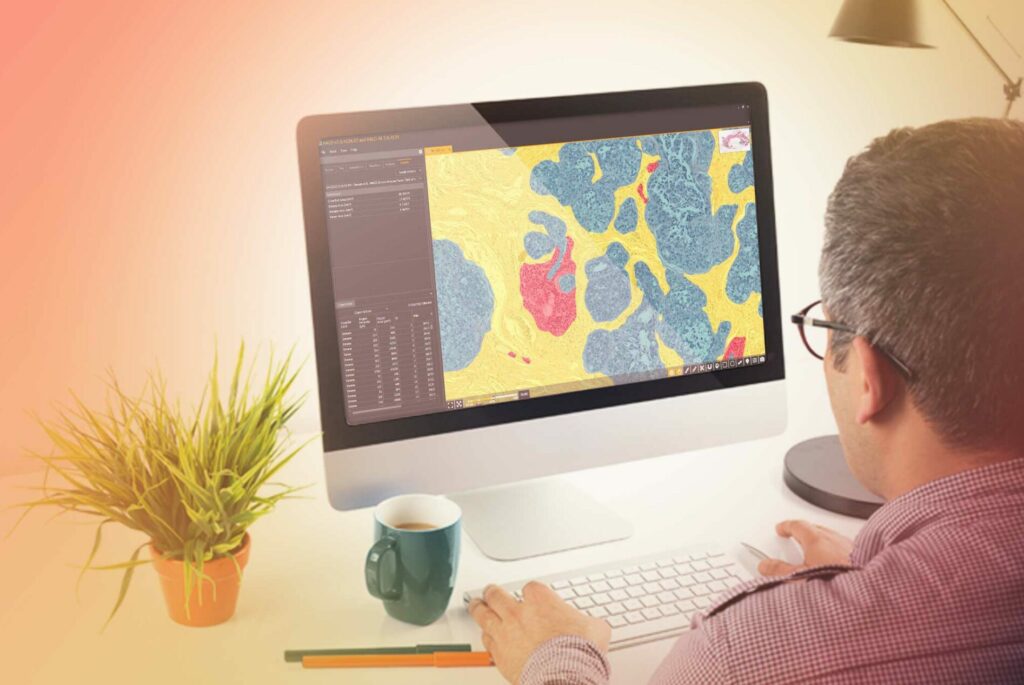
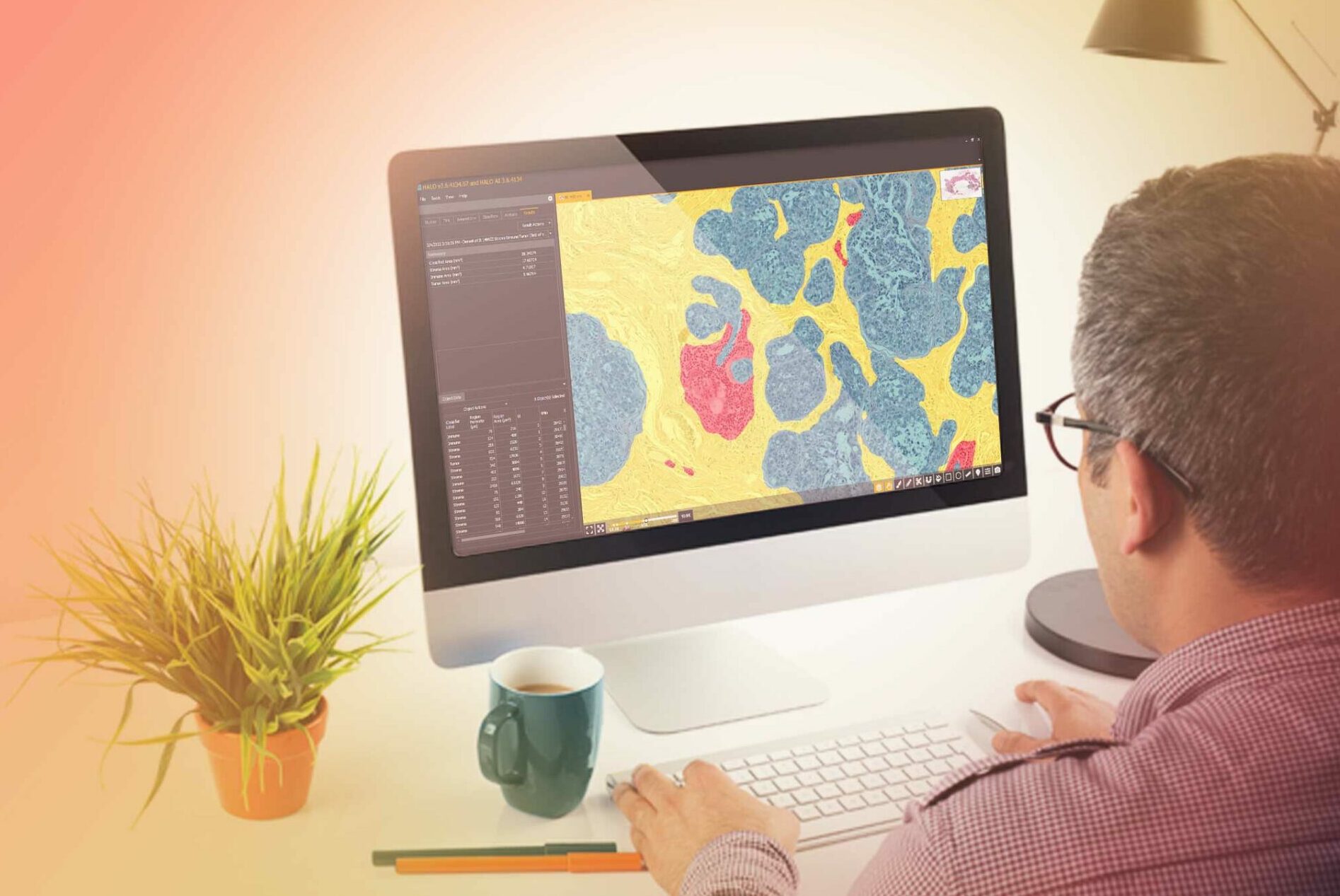
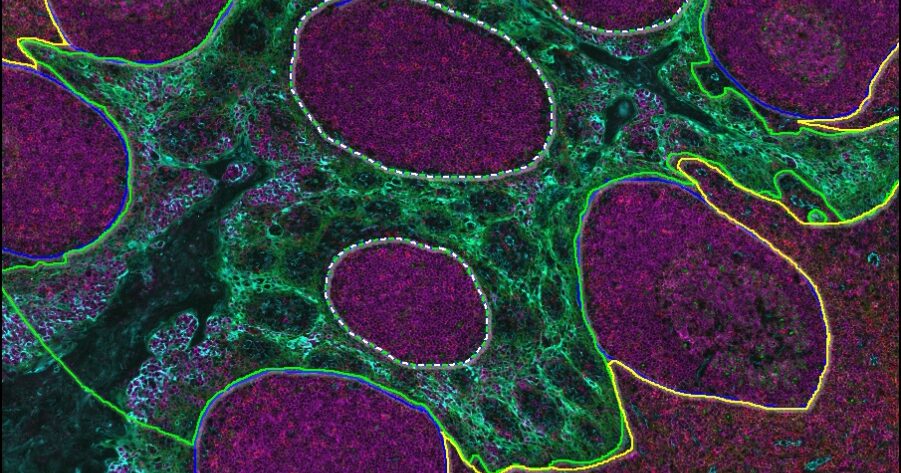
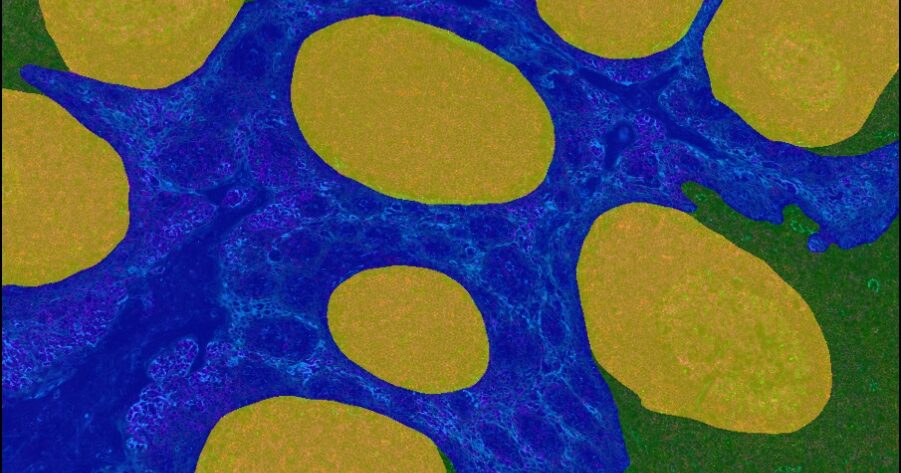
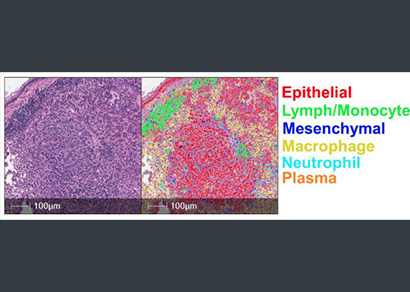
Underpinned by modern deep learning networks, HALO AI provides a collection of segmentation, classification, and phenotyping tools for brightfield and fluorescence applications. Train a classifier to quantify tissue classes, to find rare events or cells in tissues, or to categorize cell populations into specific phenotypes.
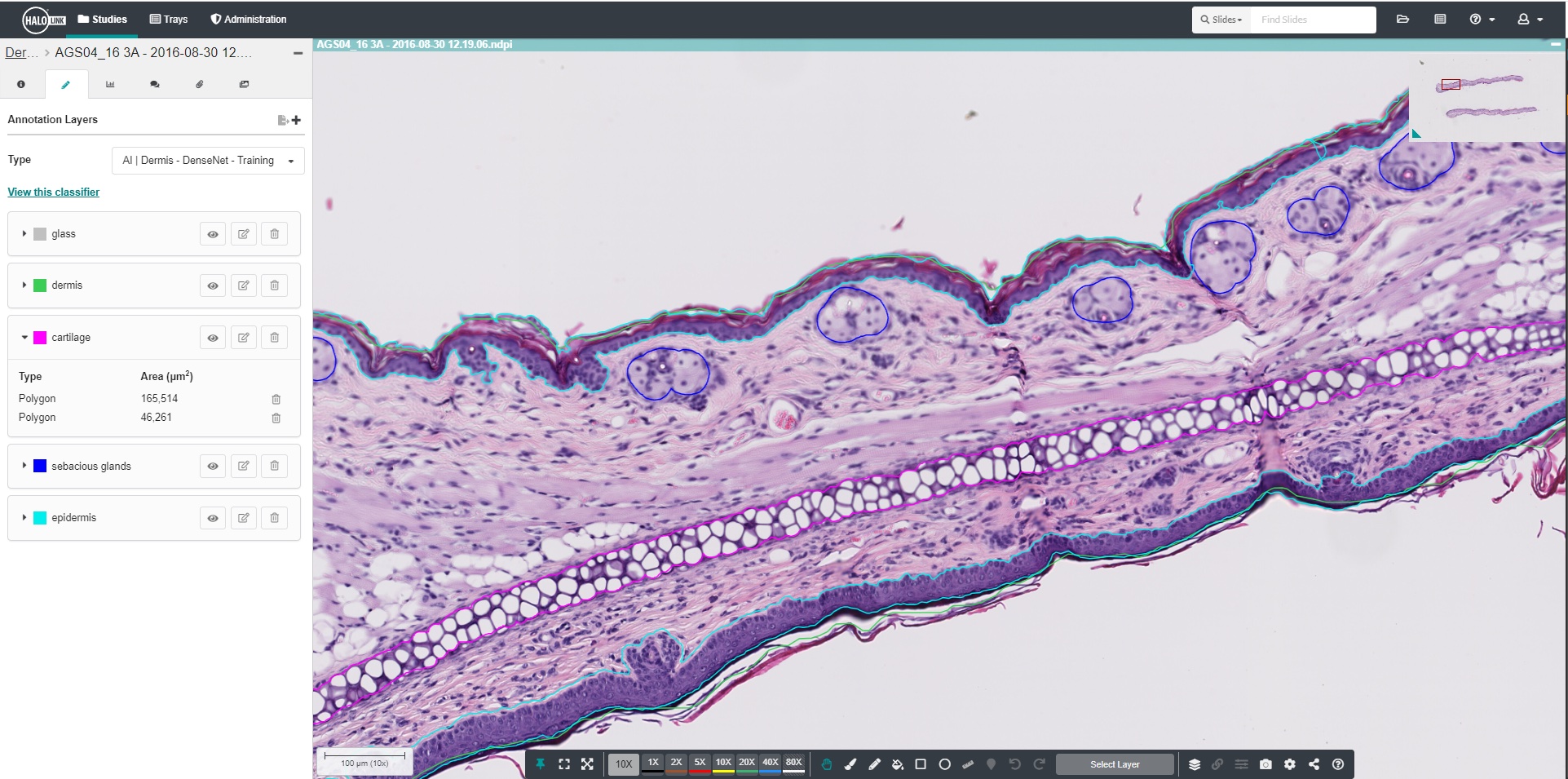
AI-Powered Annotation
Use a point and click workflow to rapidly annotate images and develop training data.
Nuclear Segmentation

Choose between several trainable AI-based nuclear segmentation networks to optimize segmentation accuracy when nuclear morphologies vary.
Membrane Segmentation

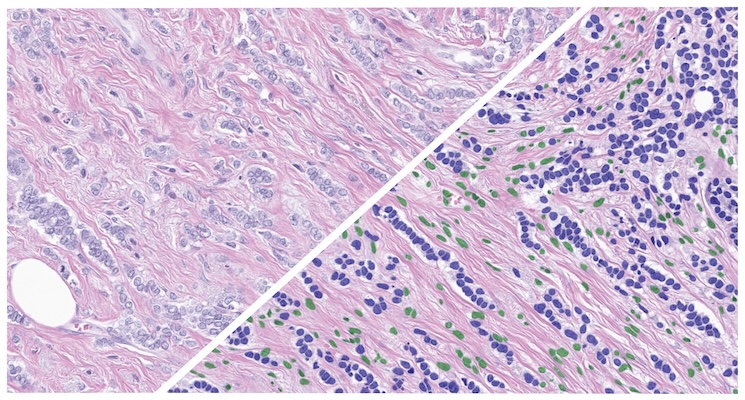
Train AI-powered membrane segmentation networks to accurately delineate cell membranes for your specific assay.
Cell & Object Phenotyping

Quickly train a phenotyper to quantify cell types or objects of interest.
Slide Quality Control

Leverage a trainable AI-powered quality control network to detect common artifacts in H&E and IHC images.
Simple & Intuitive Workflow
Choose to integrate your trained HALO AI networks in multiple locations within HALO modules to perform functions including tissue classification, nuclear and membrane segmentation, and phenotyping. Alternatively, apply HALO AI directly to an entire study of whole slide images or to select regions of interest – the choice is yours.
Manage Variability
Extreme variability runs rife in image analysis but is no match for HALO AI. Common sources of variability include diverse morphologies, alterations of morphologies from staining protocols, differences in tissue quality, uneven staining, and more.
HALO AI can be easily trained to accommodate variability to deliver accurate segmentation and classification results across large studies. HALO AI can even be trained to work across vastly different stains such as PAMS, Trichrome, H&E, and IHC.
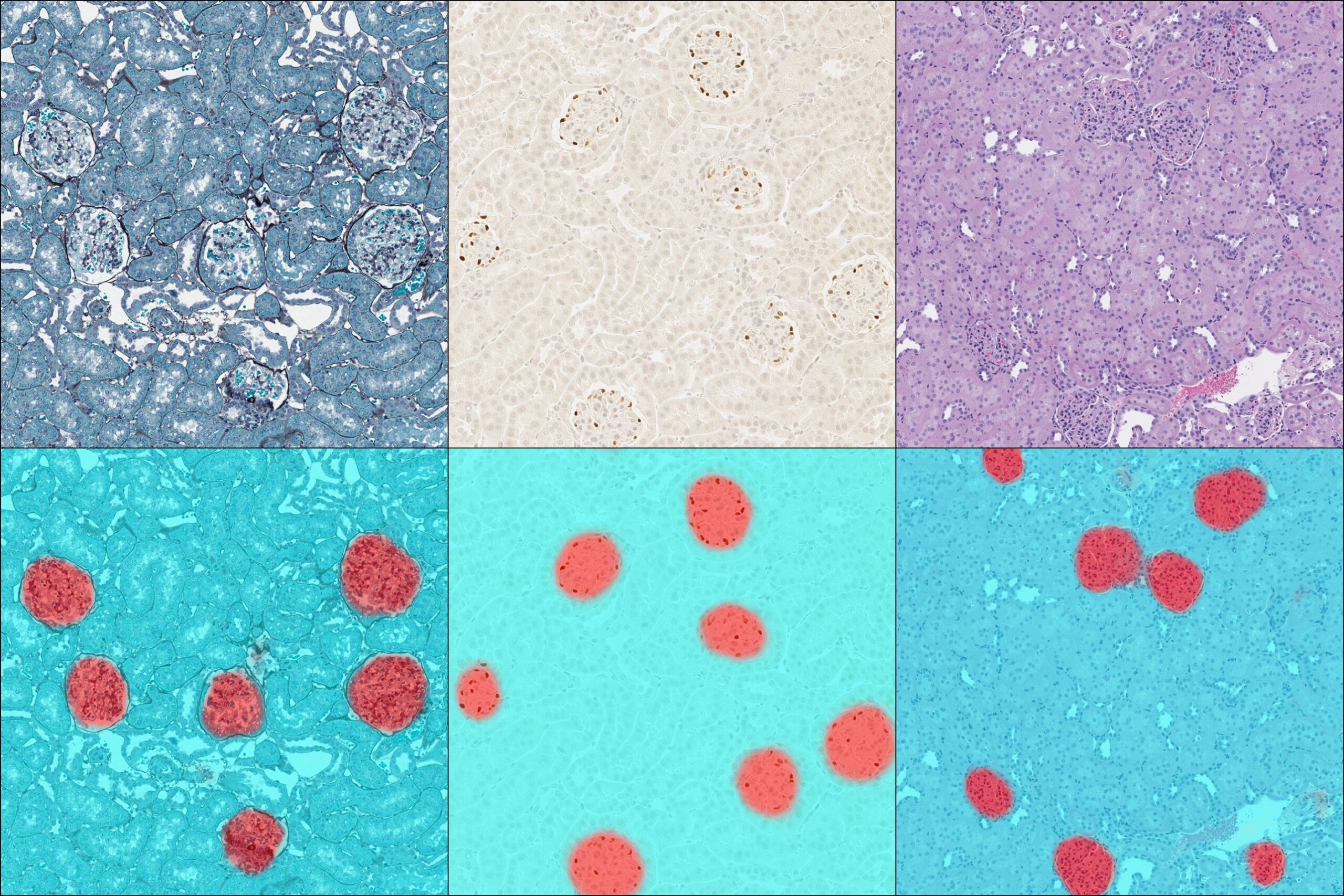
Exceptional Identification
Pre-trained nuclear and membrane segmentors available in HALO are advanced tools for nuclear and membrane segmentation, but when you need to optimize a network for a bespoke application, you need HALO AI.
As with the tissue classification and segmentation networks, you can quickly add training data with the AI-based annotation tool and train the network to optimally segment nuclei or cell membranes. And once trained, any HALO AI network can be incorporated into HALO modules for maximum utility.

Powerful Tissue Classification
The degree of variability encountered in pathology often demands an approach above and beyond a simple random forest tissue classifier.
HALO AI has two advanced pre-trained networks capable of creating high-resolution classifiers in brightfield and fluorescence, depending on how much training data and time are available for training and optimization.
Curious to see HALO AI in action on your images?
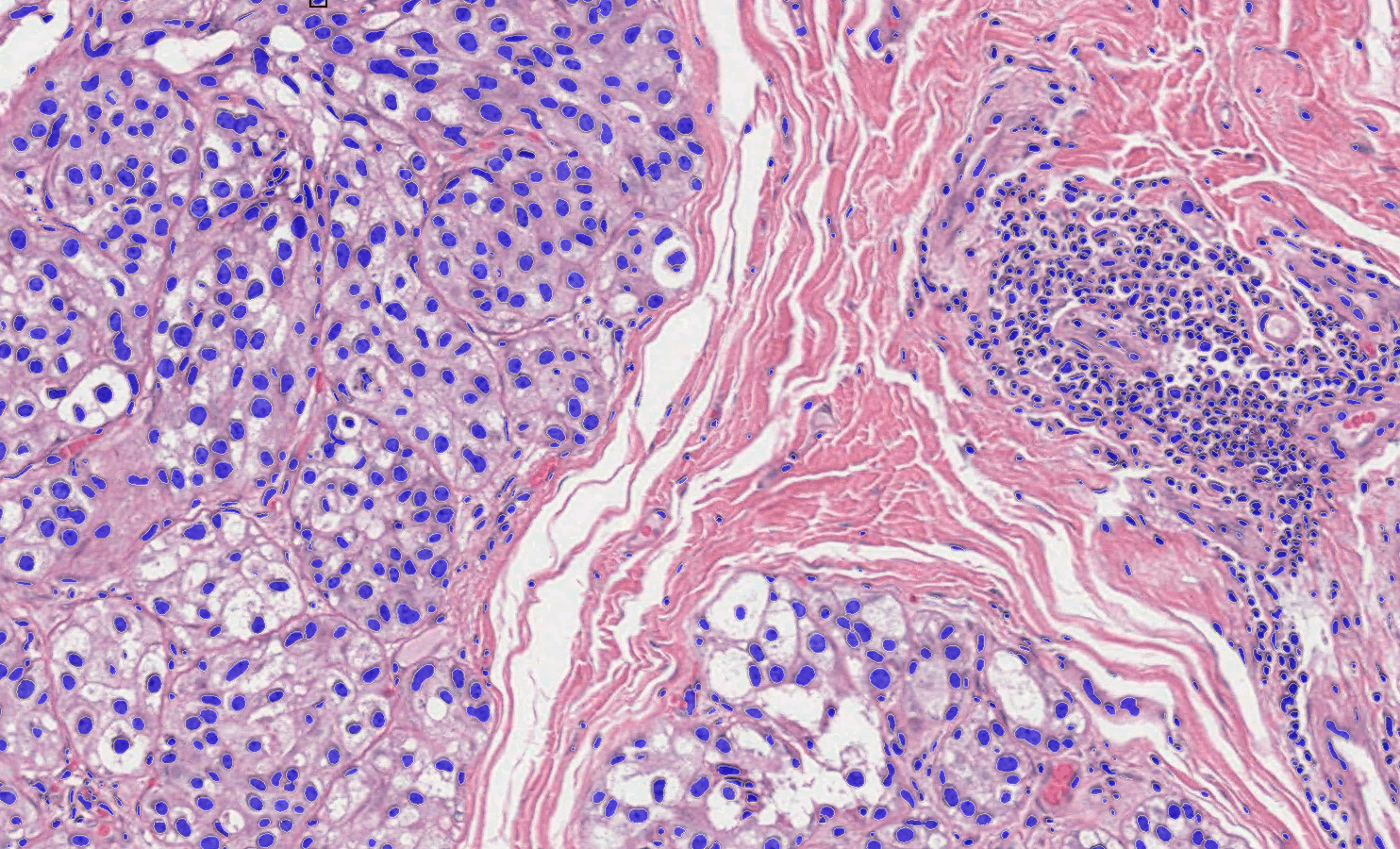
Advanced Phenotyping
Cell phenotyping can be used on any of the myriad supported image types to automatically assign cells into user defined phenotypes. Simply select a nuclear or membrane segmentation network, provide a few quick training examples, and train the network to identify phenotypes of interest.
Phenotyping is a highly effective and flexible tool to quantify cells based on morphology when biomarker information is absent and can assist with quantification and characterization of cells with complex morphologies, such as neuronal cells.
Top Notch Training and Support
At Indica Labs, we take pride in providing top-quality support. Unlimited IT support, scientific support, training, and access to our Learning Portal are included with every license at no additional cost. With support staff in the US, UK, EU, China, and Japan, you are supported in any time zone.
Wondering how easy developing your own algorithm can be without programming experience? Reach out to us to find out how intuitive HALO AI’s train-by-example interface really is.

What Our Customers Have to Say
Read independent reviews on our HALO AI Deep Learning Classifier Add-on and learn how these customers are using Indica Labs’ solutions to streamline their workflows.
Wonderful products, it is far beyond our expectations.
"Our lab both have the HALO Image Analysis System and the HALO AI, honestly speaking, the HALO system is beyond our expectations, especially the analysis process, annotation tools, the adjustable parameters, the texture and morphological recognition of nuclear and specific structure, and the after-sale care I strongly recommend you guys to choose it. it really worth the money."
lanlan Li
Panovue
Great results, can't live without this instrument!
"Scientists can achieve lots of high-quality analysis data about figures by HALO software. Many functions of HALO are very helpful to study images, such as TMA, spatial analysis, high-Plex FL, and especially HALO AI. It is easy to operate. Its interface is simple, friendly, and convenient. Once we have problems, a Halo technician is able to help us to solve them in time."
Junfeng Hao
Institute of Biophysics, Chinese Academy of Sciences
Great result. Nice tool for translational pathology research.
"We are interested in applying HALO AI in RNAscope and IHC analysis. Thanks to Yongtian ZHAO to give us wonderful training and support. We are improving and confident to apply it in the RNAscope assay development and scoring."
Fei Yang
Johnson and Johnson
Currently the best image viewer and analysis software on the market that I've used.
Nathan Aleynick
MSKCC
Platform Compatibility
HALO AI is compatible with all of the file formats that can be used in HALO and HALO Link. Yours not on the list? Email us your requirements.
- Non-proprietary (JPG, TIF, OME.TIFF) (JPG, TIF, OME.TIFF)
- Nikon (ND2)
- 3D Histech (MRXS)
- Akoya Biosciences/Quanterix (QPTIFF, component TIFF)
- Olympus / Evident (VSI)
- Hamamatsu (NDPI, NDPIS)
- Aperio/Leica Biosystems (SVS, AFI)
- Zeiss (CZI)
- Leica (SCN, LIF)
- Ventana/Roche (BIF)
- Philips (iSyntax, i2Syntax)
- KFBIO (KFB, KFBF)
- DICOM (DCM*)
*whole-slide images
Advancing Discovery with AI: Features of HALO AI
Quickly acquire training annotations on either brightfield or fluorescent images with an AI annotation tool optimized for pathology data. Simply point and click your object of interest to add an annotation.
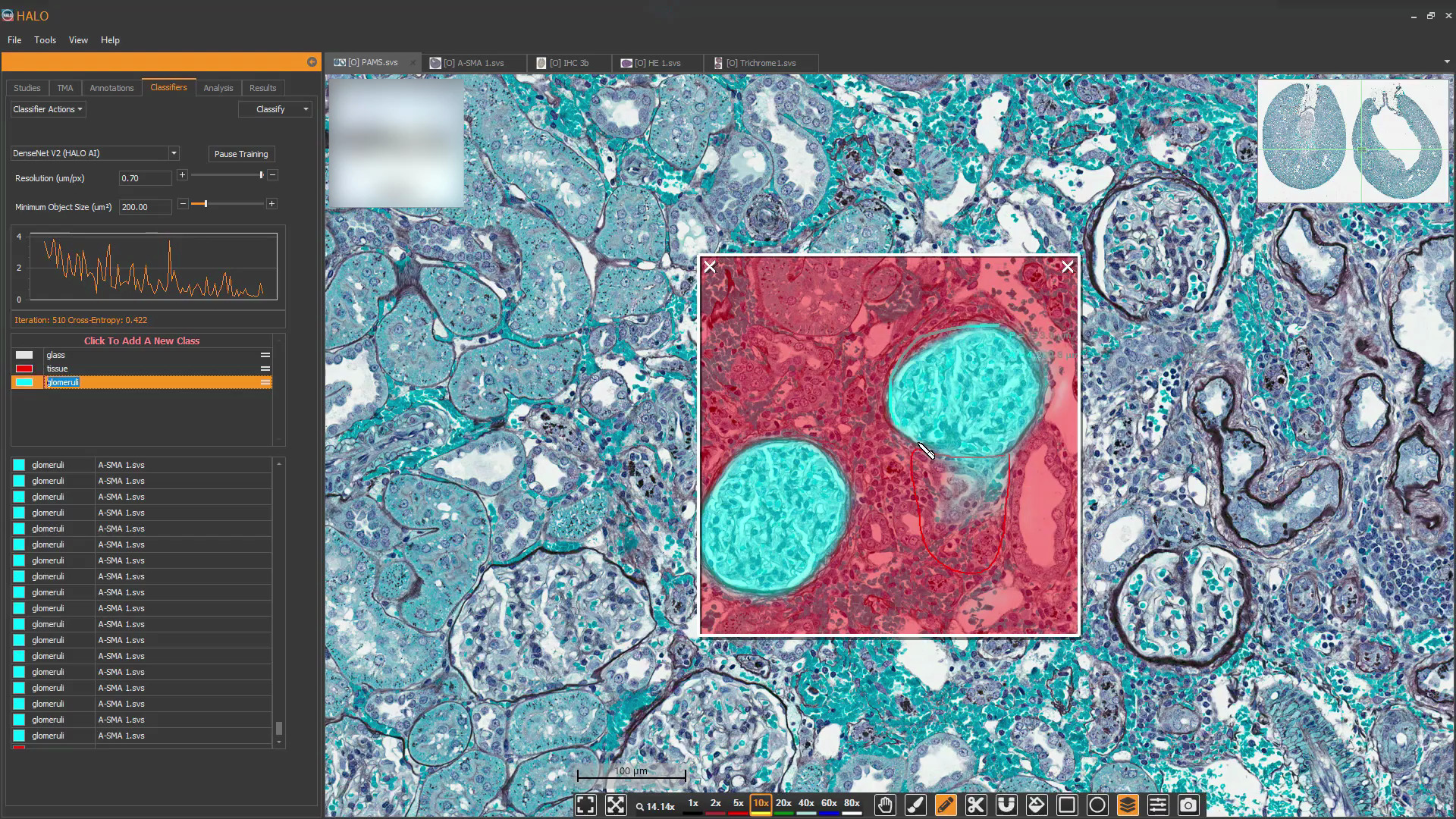
Use real-time tuning in HALO AI to watch a network as it trains in real time. Toggle the mark up on and off to evaluate performance, choose to add training data, or change parameters on-the-fly.
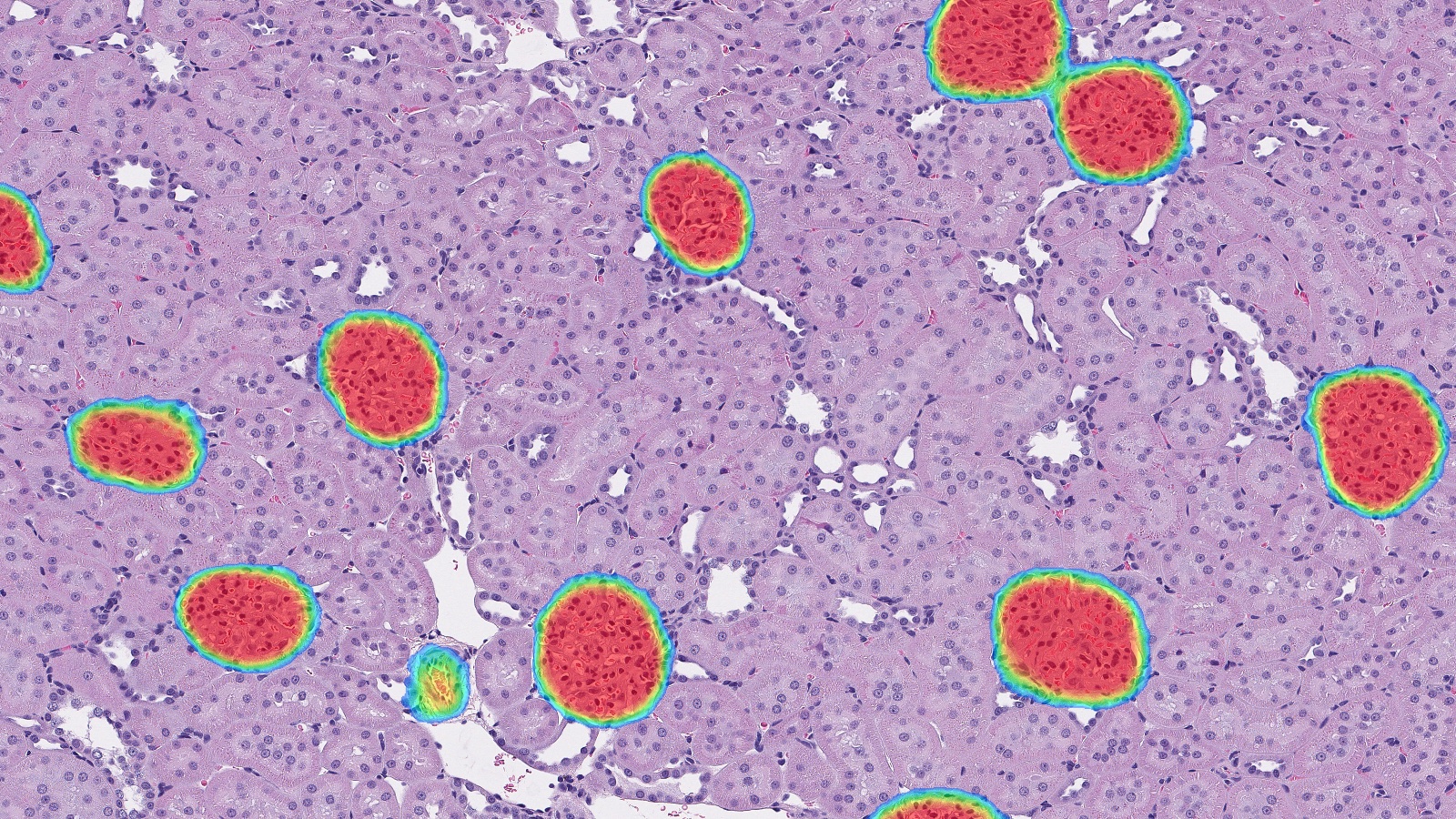
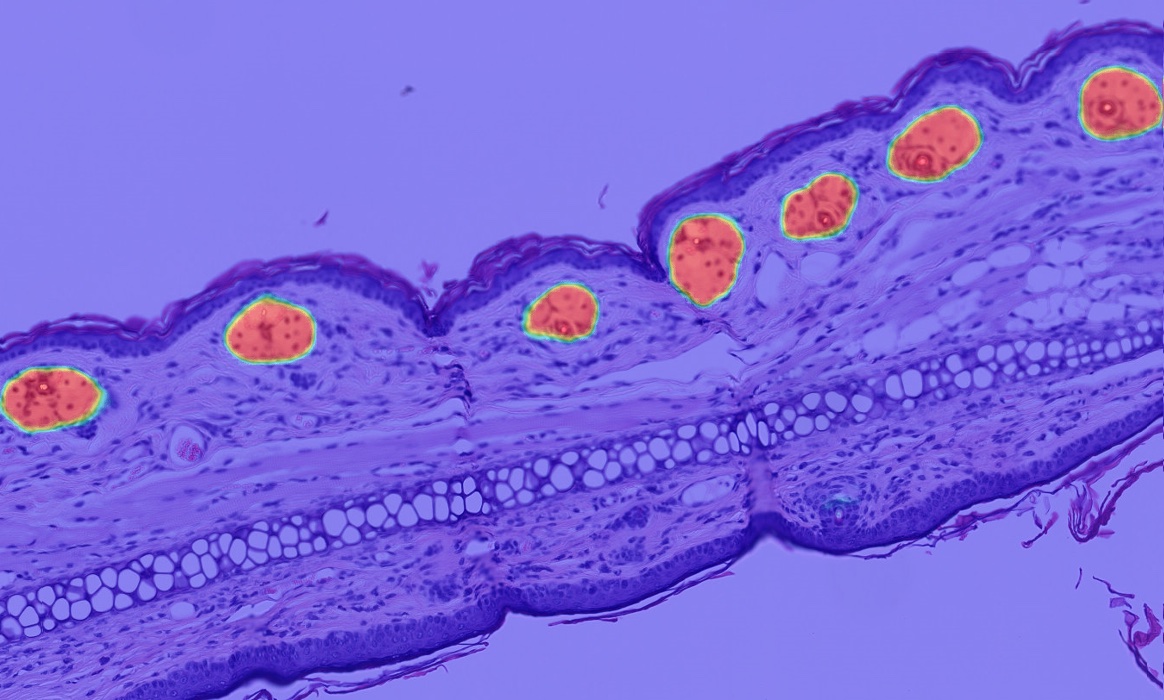
Once a HALO AI model is trained, a probability map can be used as an alternative output to a traditional mask to evaluate performance. Use real-time tuning to select an appropriate probability cut-off for a given class and view the output in a heatmap where blue represents the lowest probability and red the highest.
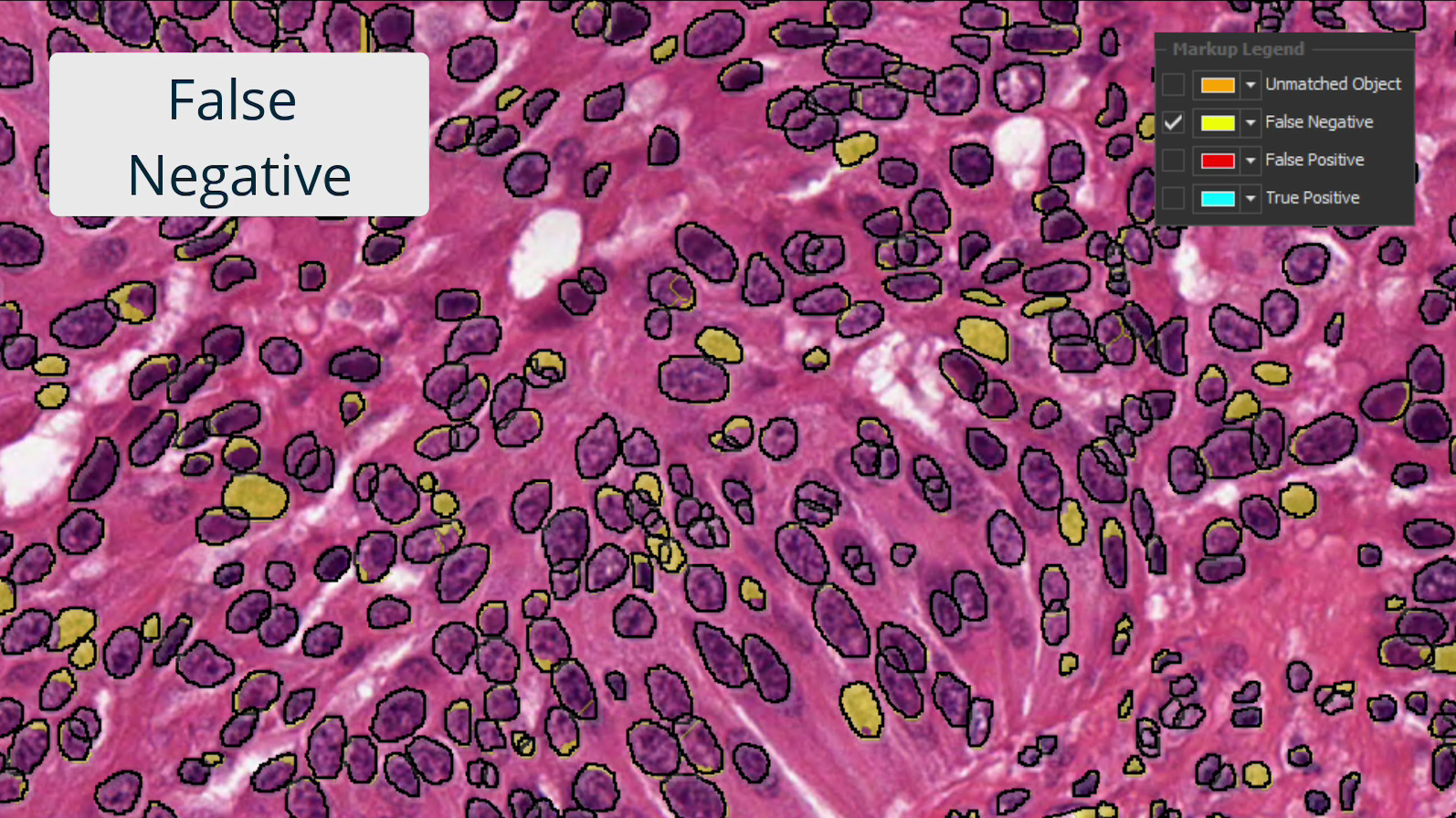
Investigate results with interactive markup images where you can toggle on and off each population of interest. Interactive markups can be combined with probability thresholding and are especially valuable in exploring validation outputs.
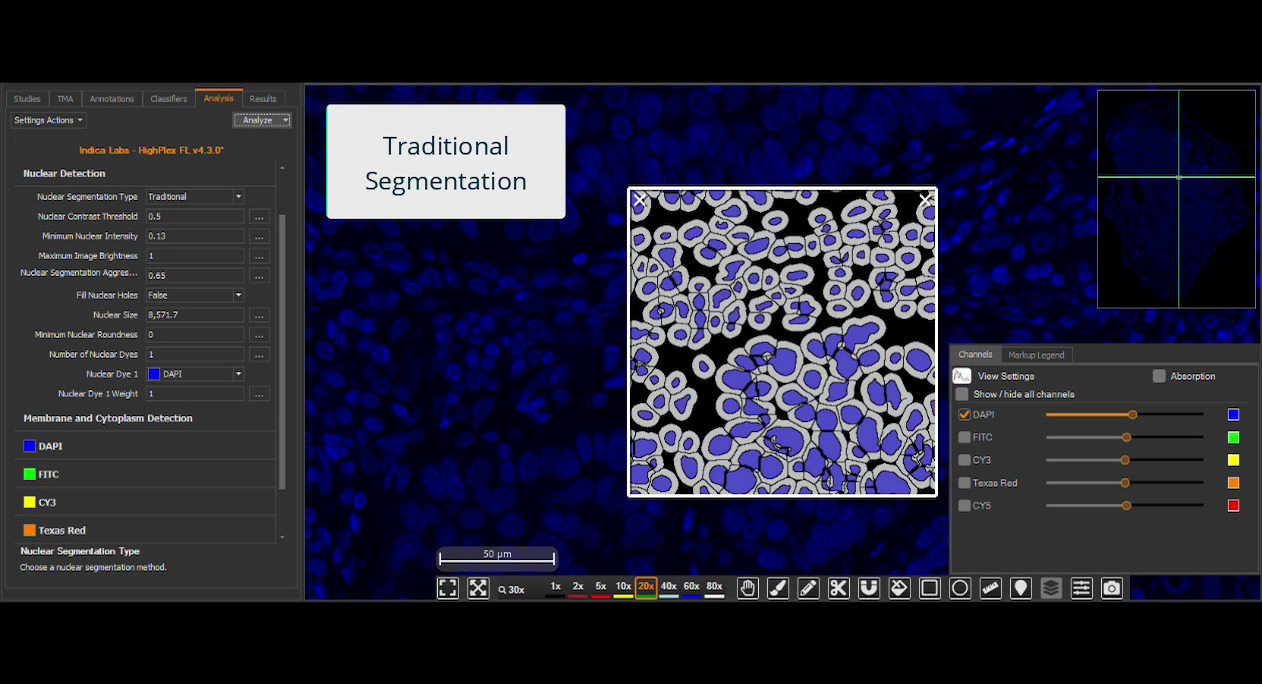
Deploy your HALO AI models in the context of HALO modules for maximum image analysis utility. Choose to utilize one or more models such as nuclear segmentation, membrane segmentation, tissue classification, cell or object phenotyping, and/or SlideQC.
Leverage the three-part Train – Validate – Test workflow to address the question: How good is my HALO AI classifier? Quantitative metrics are output based on the type of network.
Speed up your HALO AI analysis by adapting your trained network for your specific GPU. Also, you can queue training jobs based on number of iterations or time and set it to run at the end of your workday to maximize productivity and evaluate multiple model variants.
Discover HALO AI Apps
Explore our most popular HALO AI apps. You can filter by category and use the arrows at the bottom of the list to view additional options.
The Breast IHC Tumor Detection App is a pre-trained HALO AI classifier designed to detect, segment, and quantify tumor and other area across hematoxylin and DAB-stained whole-slide digital images of breast cancer.
Learn MoreThe NSCLC IHC Tumor Detection App is a pre-trained HALO AI classifier designed to detect, segment, and quantify tumor area and non-tumor area across hematoxylin and DAB-stained whole-slide digital images of NSCLC.
Learn MoreThe NSCLC IHC Cancer Cell Phenotyper App is a pre-trained HALO AI object phenotyper designed to detect, segment, and quantify non-cancer cells, IHC-positive cancer cells and IHC-negative cancer cells across hematoxylin and DAB-stained whole-slide digital images of NSCLC.
Learn MoreThe Breast IHC Cancer Cell Phenotyper App is a pre-trained HALO AI object phenotyper designed to detect, segment, and quantify cancer cells and other cells across hematoxylin and DAB-stained whole-slide digital images of breast cancer.
Learn MoreThe NSCLC H&E Cancer Cell Phenotyper App is a pre-trained object phenotyper designed to detect, quantify, and segment cancer cells from non-cancer cells across H&E-stained whole-slide digital images of NSCLC.
Learn MoreThe CRC H&E Cancer Cell Phenotyper App is a pre-trained HALO AI object phenotyper designed to detect, segment, and quantify cancer cells and other cells across H&E-stained whole-slide digital images of colorectal cancer.
Learn MoreThe Pan-Cancer H&E Lymphocyte Cell Phenotyper App is a pre-trained HALO AI object phenotyper designed to detect and quantify lymphocytes across whole slide H&E-stained images of multiple tumor types.
Learn MoreThe Gastric H&E Tumor Tissue Detection App is a pre-trained HALO AI masking classifier designed to segment tumor, stroma, necrosis/other, and glass areas across H&E-stained whole slide images of gastric cancer.
Learn MoreThe Head & Neck Squamous Cell Carcinoma (HNSCC) H&E Tumor Tissue Detection App is a pre-trained HALO AI masking classifier designed to segment tumor, stroma, necrosis/other, and glass area across H&E-stained whole slide HNSCC images.
Learn MoreThe Non-Small Cell Lung Cancer (NSCLC) H&E Tumor Tissue Detection App is a pre-trained HALO AI masking classifier designed to segment tumor, stroma, necrosis/other, and glass area across H&E-stained whole slide images of NSCLC.
Learn MoreThe Ovarian H&E Tumor Tissue Detection App is a pre-trained HALO AI masking classifier designed to segment tumor, stroma, necrosis/other, and glass area across H&E-stained whole slide images of ovarian cancer.
Learn MoreThe Breast H&E Cancer Cell Phenotyper App is a pre-trained HALO AI phenotyper designed to detect and quantify cancer cells across whole slide H&E-stained images of breast cancer tissue.
Learn MoreFree Image Analysis
See HALO AI in action on up to three of your images with a free analysis.
Knowledge Center
Learn more about HALO AI by exploring the tabs below. For an introduction to the capabilities of HALO AI, we recommend you check out our HALO AI white paper!

Characterizing the TME using IMC and HALO® Image Analysis
See in this app note how automated analysis of highly multiplexed IMC images using HALO and HALO AI yields rich cellular and spatial data from a streamlined workflow.

COMET™ and RNAscope™ Image Analysis Using HALO and HALO AI
Read this collaborative app note to learn how our Pharma Services team leveraged the HALO FISH-IF and Spatial Analysis modules and HALO AI for analysis of COMET™ and RNAscope™ images.

Glomeruli Quantification and Characterization Application Note
Don’t let varied stain selections hold back your digital image analysis of kidney sections! With HALO AI you can develop a tissue classifier that accurately detects and segments glomeruli in sections across a range of stains including H&E, trichrome, PAMS, and DAB with hematoxylin.

HiPlex RNAscope™ Application Note
Using our FISH module we demonstrate how to perform 12-plex RNAscope image analysis using ACD’s HiPlexv2 assay.

RNAscope Image Analysis Using HALO and HALO AI
Using our ISH module for brightfield and our FISH module for fluorescence, we demonstrate how to quantify ISH signal from RNAscope® assays on a per cell or per unit area basis and output an overall expression score based on ACD Bio’s recommended scoring guidelines.

HALO Platforms for Life Sciences Brochure
Download our brochure to learn about our HALO, HALO AI, and HALO Link platforms, and how they can advance your digital pathology image analysis and management workflows.

HALO, HALO AI, & HALO Link Cloud Services Brochure
Discover how our Cloud Services team can provide a secure, scalable, and flexible cloud deployment of HALO, HALO AI, and HALO Link.

Cloud Services and AI Diagnostics Case Study
Learn how the optimized, scalable infrastructure provided by Indica Labs Cloud Services enables the rapid development and robust collaboration of our AI Diagnostics group, powering the next generation of automated AI-based decision support tools.

HALO in the Cloud at the University of Washington
Read this case study to learn how partnering with our Cloud Services team has delivered greater scalability and cost efficiency for the BioRepository and Integrated Neuropathology laboratory at University of Washington Medicine.

NCI Case Study
Download this case study to learn about considerations the National Cancer Institute made when planning to migrate its HALO, HALO AI, and HALO Link deployments to the cloud, and benefits the NCI experienced after migration, including 50% faster HALO image analysis.

Clustering Workflow eBook
Check out our collaborative eBook with Lunaphore to learn about an optimized workflow for extracting information from hyperplex datasets and illuminating changes in cellular neighborhoods using HALO, HALO AI, and the Lunaphore COMET platform.

Lunaphore COMET Hyperplex IF and HALO® Image Analysis eBook
Our collaborative eBook with Lunaphore shows how HALO and HALO AI, alongside the Lunaphore COMET instrument, enable a high-throughput workflow for phenotypic and spatial analysis of cells in the tumor microenvironment.

Qualitative and Quantitative Evaluation of the TME eBook
Check out this ebook to see how AI-based nuclear segmentation and tissue classification can streamline cell phenotyping across tissue compartments and samples.

Sequential Same Slide mIF and H&E eBook
Reveal more data on a single slide with this workflow for sequential same slide multiplex IF and H&E staining and imaging using HALO and HALO AI.

AI-Assisted Quality Control for Artifact Detection: Deployment in Image Management System with GLP Support
Download our poster to learn how artifact detection with the AI-powered SlideQC BF can advance your quality control pipelines with the GLP support of HALO Link.
All-In-One Digital Pathology: Compare and Contrast the Tumor Microenvironment of Renal Cell Carcinoma Tissue with Paired, Patient Derived Tumoroids
Check out this collaborative poster to learn about a streamlined triplex chromogenic workflow leveraging HALO and HALO AI for analysis of slides assayed with Leica Biosystems’ ChromoPlex III Triple Detection RUO.

An Automated Deep Learning Artifact Detection Tool for Quality Control of Whole-Slide Digital Pathology Images
Explore SlideQC: an AI-based tool which identifies pre-analytic processing errors in whole-slide images for quality control in digital pathology laboratories.
Automated Tumor Budding Quantification in Colorectal Carcinoma H&E Images
Learn about quantification of tumor budding in colorectal carcinoma using HALO and HALO AI.
Automated, Flexible Multiplex Immunofluorescence for Tumor Microenvironment Profiling Using HCR Gold IF and Clinically-Relevant Antibody Clones
Check out this collaborative poster to learn about a streamlined triplex chromogenic workflow leveraging HALO and HALO AI for analysis of slides assayed with Leica Biosystems’ ChromoPlex III Triple Detection RUO.
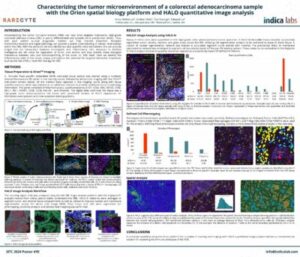
Characterizing the TME of a Colorectal Adenocarcinoma Sample with the Orion Spatial Biology Platform and HALO Image Analysis
Download our collaborative poster with RareCyte to learn how to analyze a colorectal adenocarcinoma sample with HALO and HALO AI
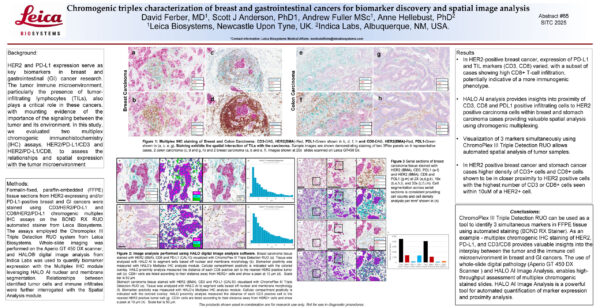
Chromogenic triplex characterization of breast and gastrointestinal cancers
Check out this collaborative poster to learn about a streamlined triplex chromogenic workflow leveraging HALO and HALO AI for analysis of slides assayed with Leica Biosystems’ ChromoPlex III Triple Detection RUO.
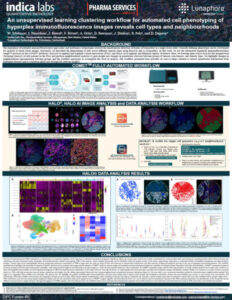
Clustering Workflow for Advanced Cell Phenotyping of Hyperplex IF Images
Download our collaborative poster with Lunaphore to learn about a clustering workflow that uses unsupervised learning for COMET™ images.
Our HALO AI customers are making vital discoveries in oncology, neuroscience, and metabolism research with over 150 publications and growing! Check out some of the research our customers are publishing.
HALO AI High Impact Publications
Review this selection of high-impact publications for examples of HALO AI applications in fields ranging from metabolism to immuno-oncology and myology.

HALO AI Publications and Creative Applications
Check out this publication list to find recent high impact HALO AI publications and highlights of several studies leveraging AI in creative or unusual applications.

HALO Image Analysis in Drug Discovery
Review this publication list to learn how HALO platforms are advancing image analysis and helping accelerate findings throughout the drug discovery and development pipeline.

HALO AI Publications and Creative Applications
Check out this publication list to find recent high impact HALO AI publications and highlights of several studies leveraging AI in creative or unusual applications.
Submit the form below to view the requested document

HALO AI High Impact Publications
Review this selection of high-impact publications for examples of HALO AI applications in fields ranging from metabolism to immuno-oncology and myology.
Submit the form below to view the requested document
Fetal growth restriction (FGR) is a significant concern in obstetrics, affecting 5–10% of pregnancies globally. FGR is a major contributor to perinatal morbidity and mortality and poses a significant maternal risk due to potential co-occurrence with pre-eclampsia. Despite their impact,...
Learn MoreThis study examined CD8+ cell distribution in hepatocellular carcinoma (HCC) and peritumoral liver tissue to investigate the overall survival (OS) and recurrence-free survival (RFS). It is well understood that CD8+ lymphocytes are involved in both the anti-tumor response and in...
Learn MoreEZH2 is the catalytic component of Polycomb Repressive Complex 2 (PRC2) and performs trimethylation of histone H3 at lysine 27 (H3K27me3) to silence chromatin. PRC2 is known to play different roles in different cancers and inhibitors of PRC2 histone methyltransferase...
Learn More
HALO AI Whitepaper
Check out this white paper for an in-depth introduction to the deep learning-based add-on to HALO, including available networks, common functions, and the “train – validate – test” Validation workflow.

Pharma Services and HALO Link White Paper
Download our white paper to learn how Indica Labs’ Pharma Services collaborates with customers and delivers HALO and HALO AI image analysis using HALO Link.

Quantitative Multiplex Chromogenic Image Analysis Guide
From stain and chromogen selection to honing nuclear and membrane detection in HALO, this comprehensive guide provides best practices and optimization techniques to help take your multiplex chromogenic image analysis to the next level.

Quantitative RNAscope™ Image Analysis Guide
From experimental design considerations to optimized setup of HALO image analysis parameters, our guide will help take your quantitative RNAscope™ image analysis to the next level.
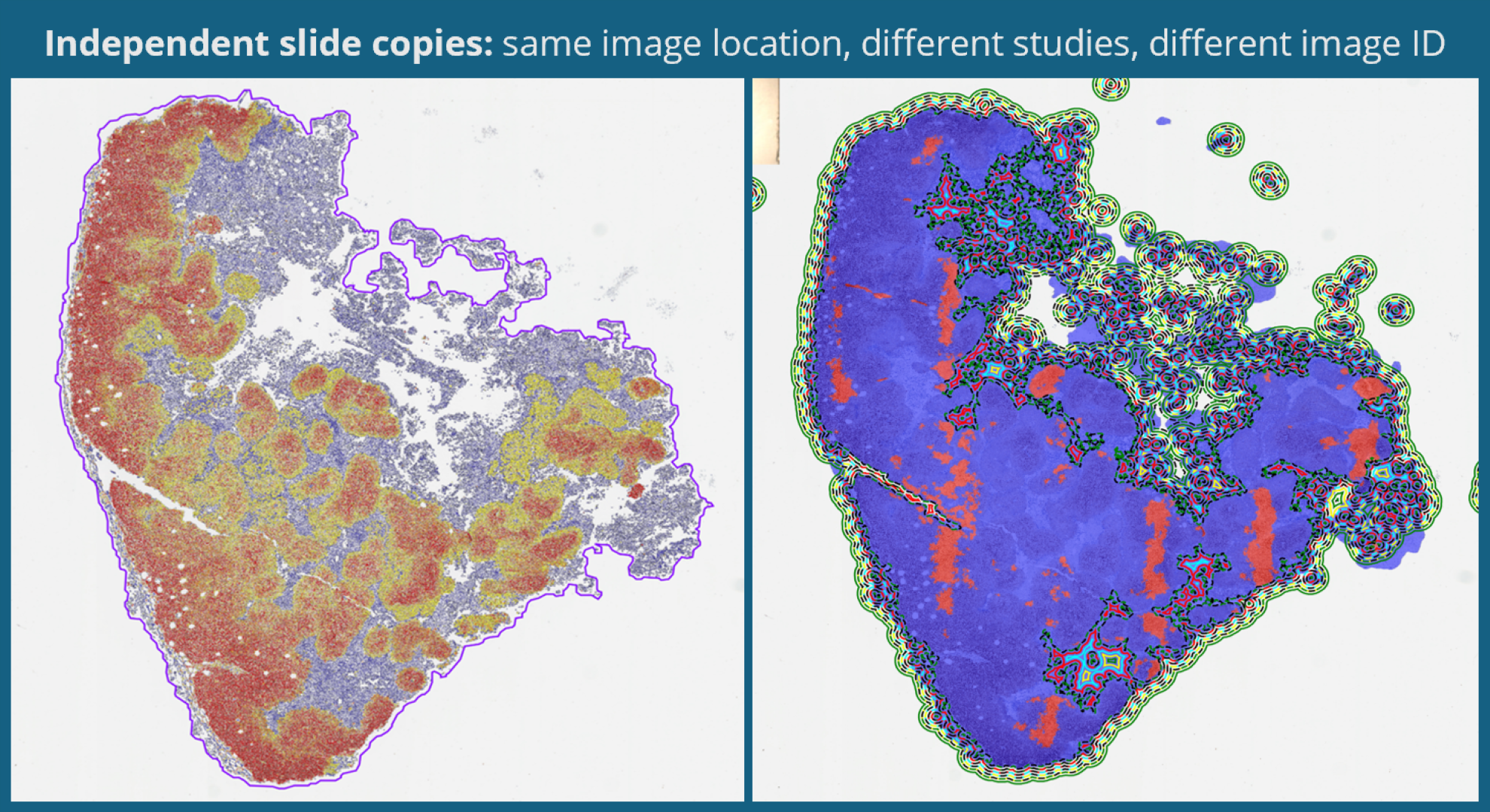

What is new in HALO®, HALO AI, and HALO Link 4.3?
11 June 2026 | Learn about the exciting new features in the 4.3 versions of HALO, HALO AI, and HALO Link!
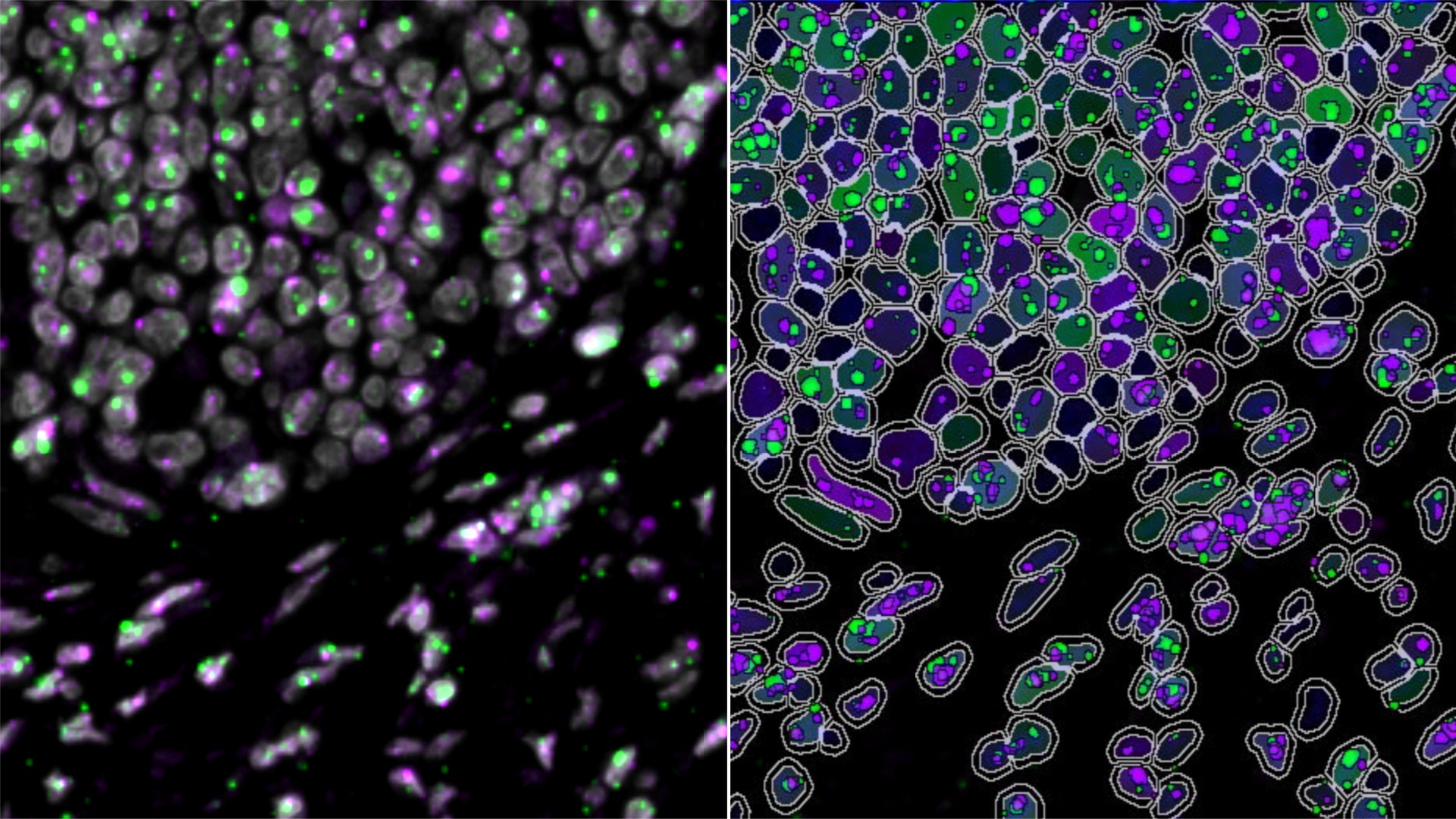
From Images to Insights: Getting Started with RNAscope™ Image Analysis in HALO®
25 June 2026 | Discover commonly used tools to get started today with RNAscope™ image analysis in HALO®
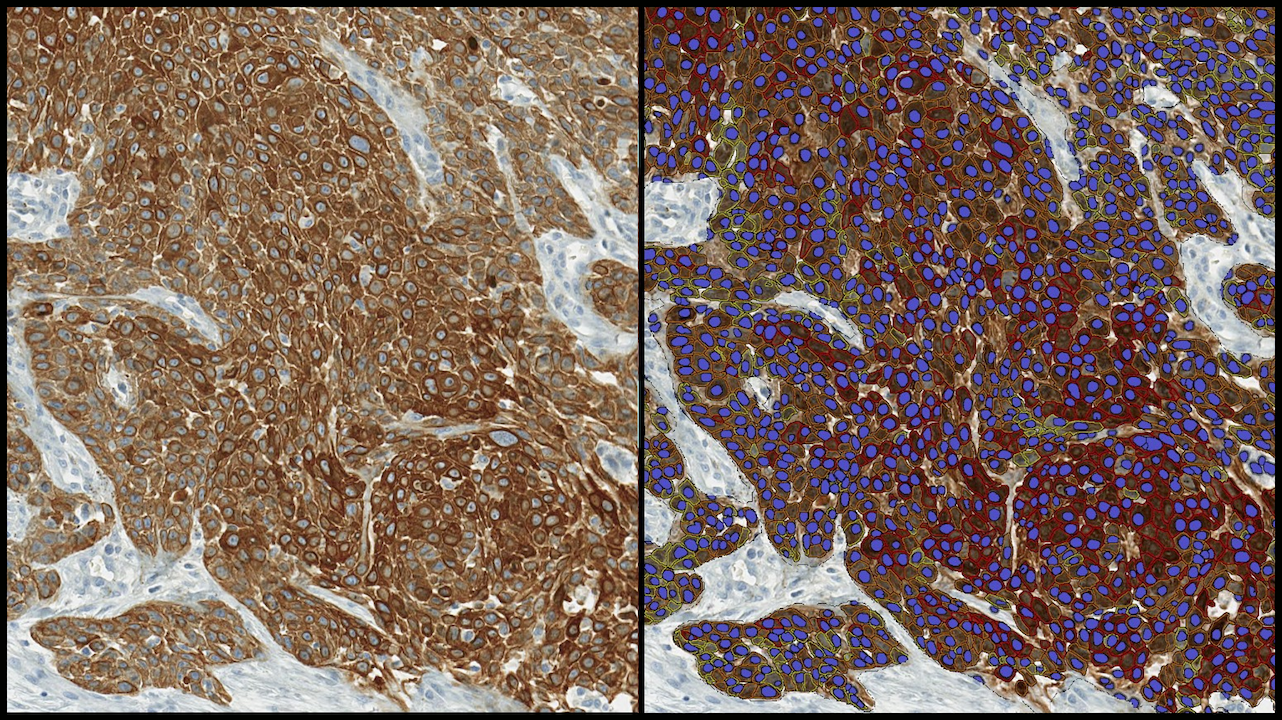
Getting Started with AI-Powered HALO Image Analysis
23 April 2026 | Learn how HALO and HALO AI enable quantitative image analysis, turning complex tissue data into actionable insights
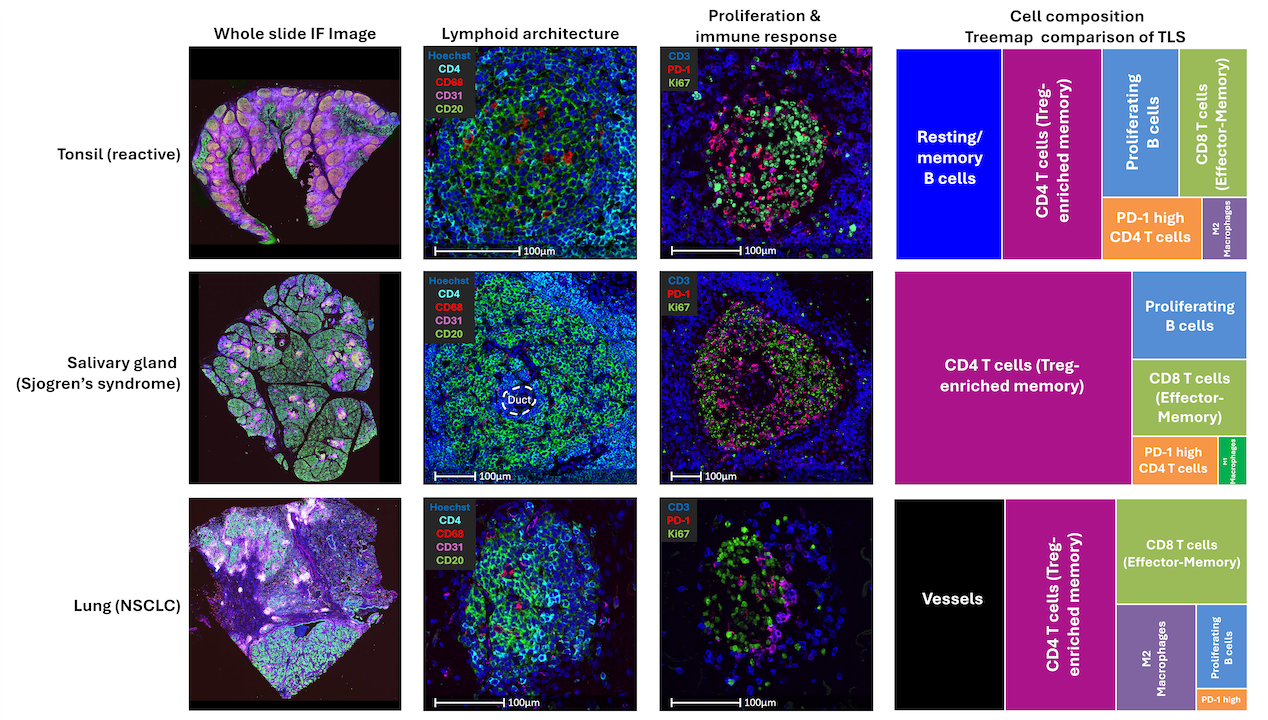
Masterclass: Unlock Cell Population Discovery Using HALO® High Dimensional Analysis
31 March 2026 | Learn how unbiased high-dimensional analysis can be seamlessly combined with AI-powered HALO workflows for multiplex images!
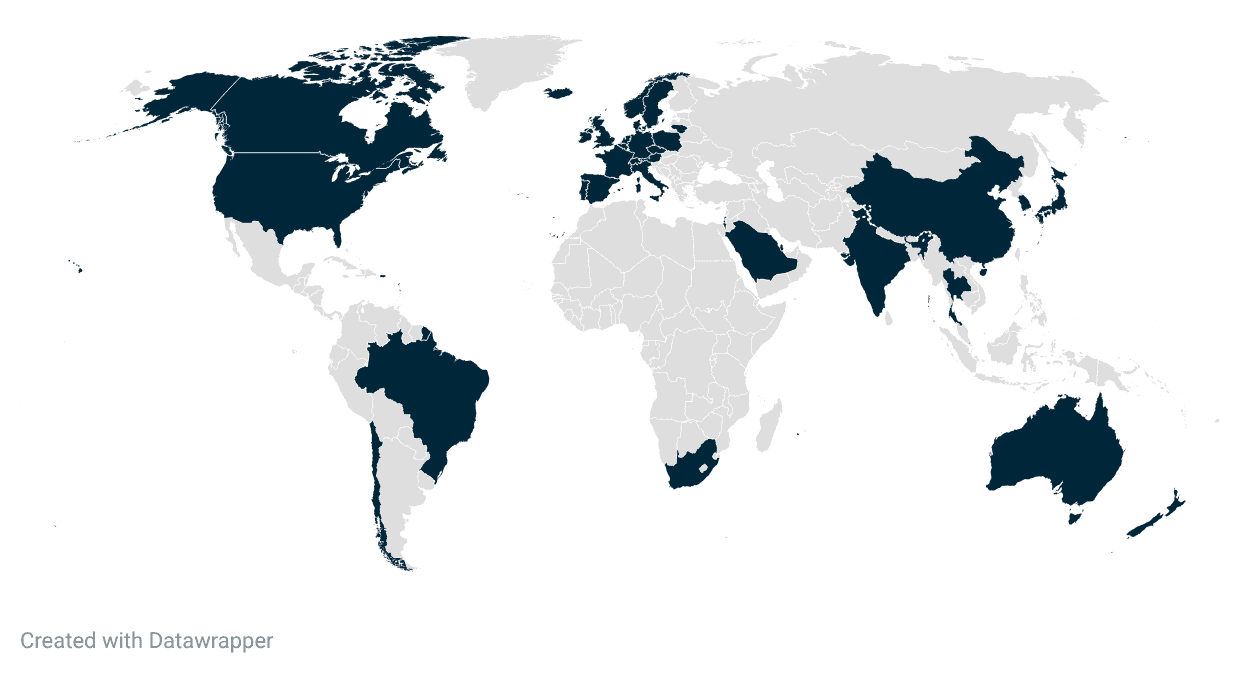
Indica Labs Surpasses 3,000 Deployments of HALO Digital Pathology Software Worldwide Since 2013
Albuquerque, NM, March 3, 2026 – Indica Labs, the global leader in AI-powered digital pathology software and services, today announces surpassing the 3,000th installation of

Indica Labs’ Pharma Services Launches GCP-Compliant Image Analysis Service
Albuquerque, NM, July 8, 2025 – Indica Labs, the global leader in AI-powered digital pathology software and services, today announces that their Pharma Services team

Indica Labs Joins the AWS Partner Network
Albuquerque, NM, December 4, 2024 – Indica Labs, the global leader in AI-powered digital pathology software and services, announced today that they have joined the

The Institute of Molecular Pathology and Immunology of the University of Porto (IPATIMUP) Selects Indica Labs’ HALO AP® to Deliver AI-enabled Diagnostics
Albuquerque, NM, October 30, 2024 – The Institute of Molecular Pathology and Immunology of the University of Porto (IPATIMUP), among the leading institutes for clinical

Looking Back: Reviewing 2025 at Indica Labs
As we reach the end of 2025, the Indica Labs team extends our sincere thanks to the customers and collaborators who helped make this year
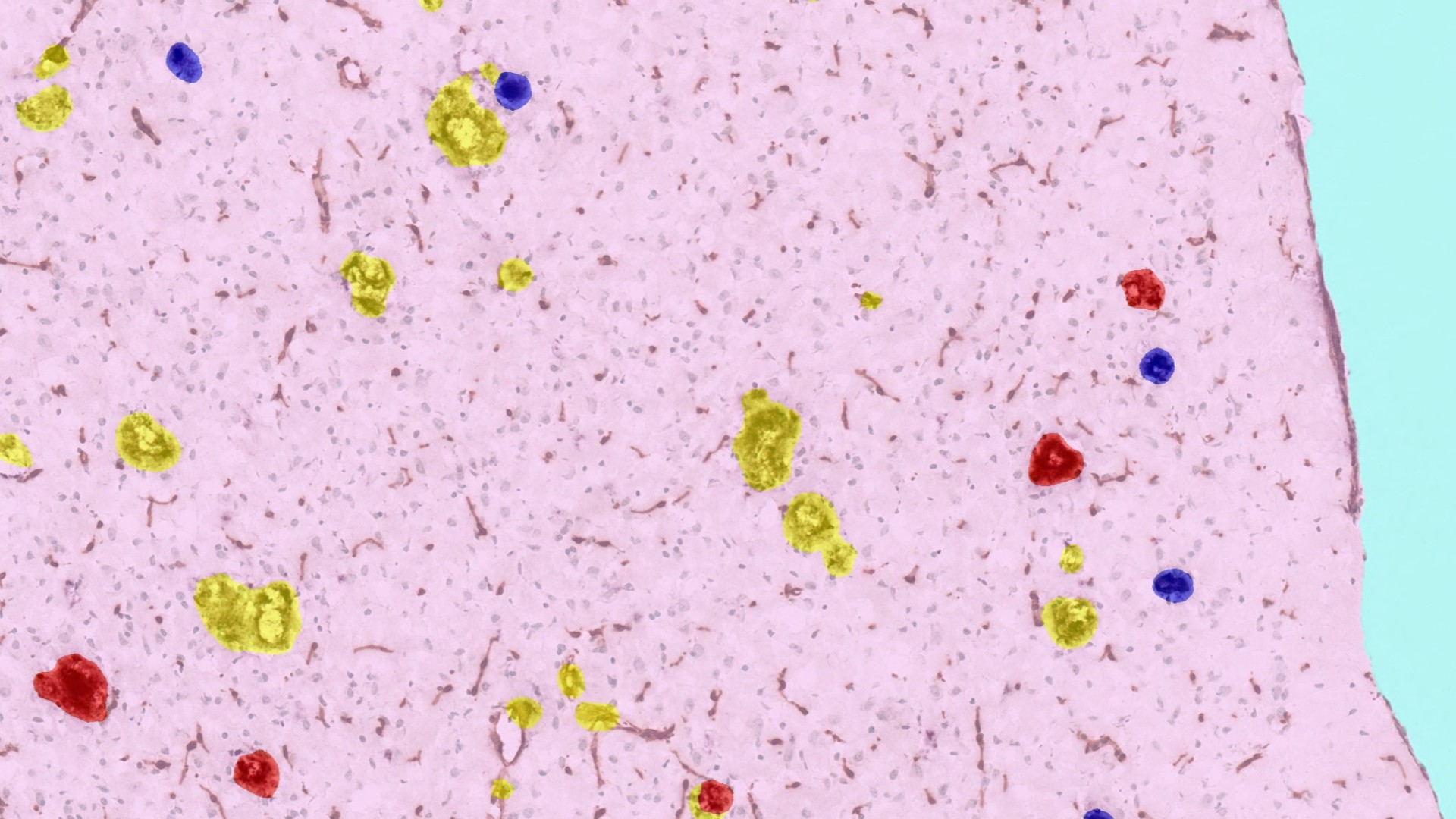
Advancing Neuroscience Research with HALO® and HALO AI
As scientists work to continue unraveling the structural, cellular, and molecular complexities of the brain and its diseases, the characteristics of this unique tissue pose
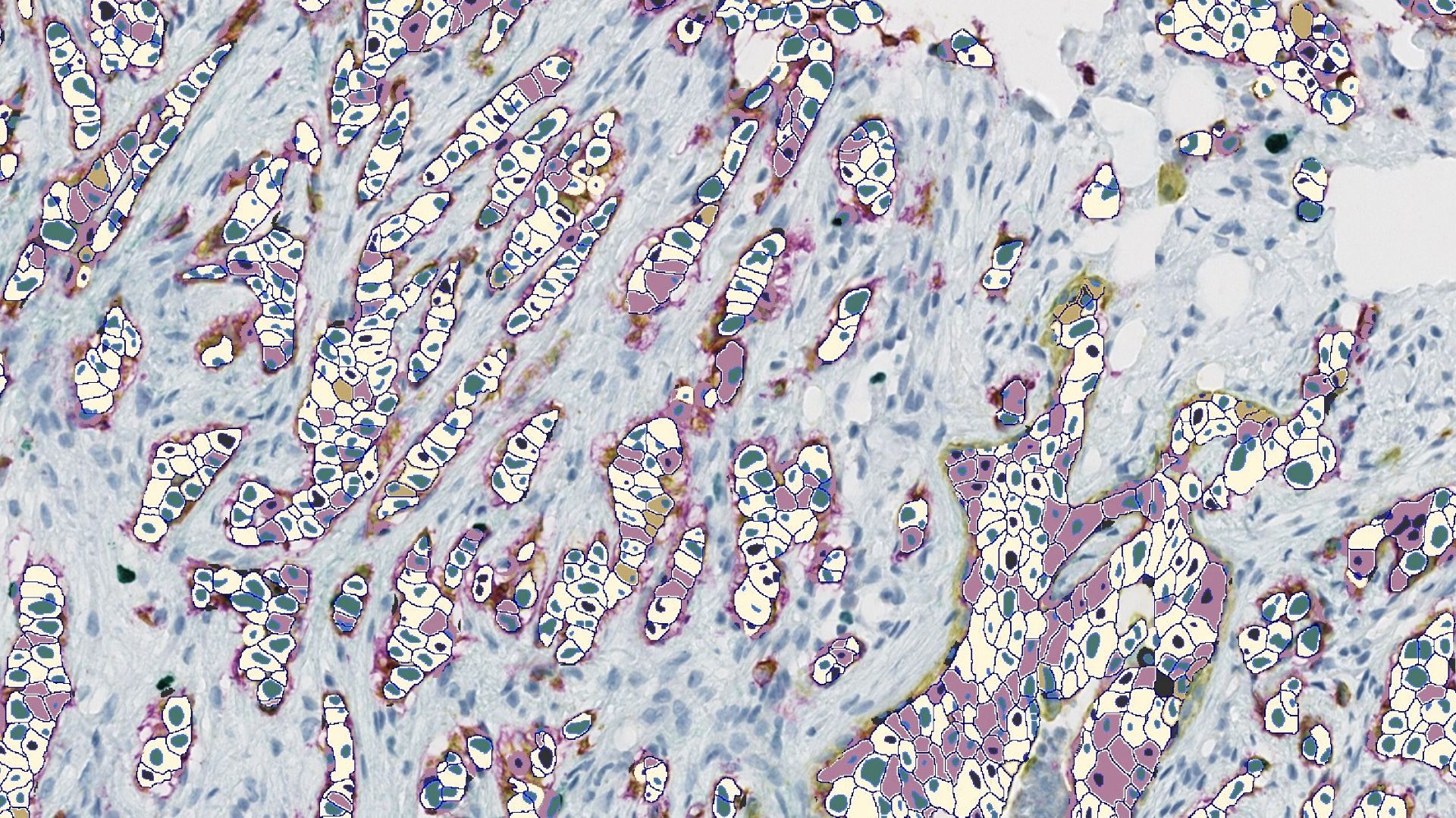
Unlocking the Potential of Antibody-Drug Conjugates (ADC) with HALO® Image Analysis and Indica Labs Pharma Services
Antibody-drug conjugates (ADC) represent one of the most promising classes of next-generation cancer therapeutics, combining antibody-based targeting with the potency of chemotherapy.
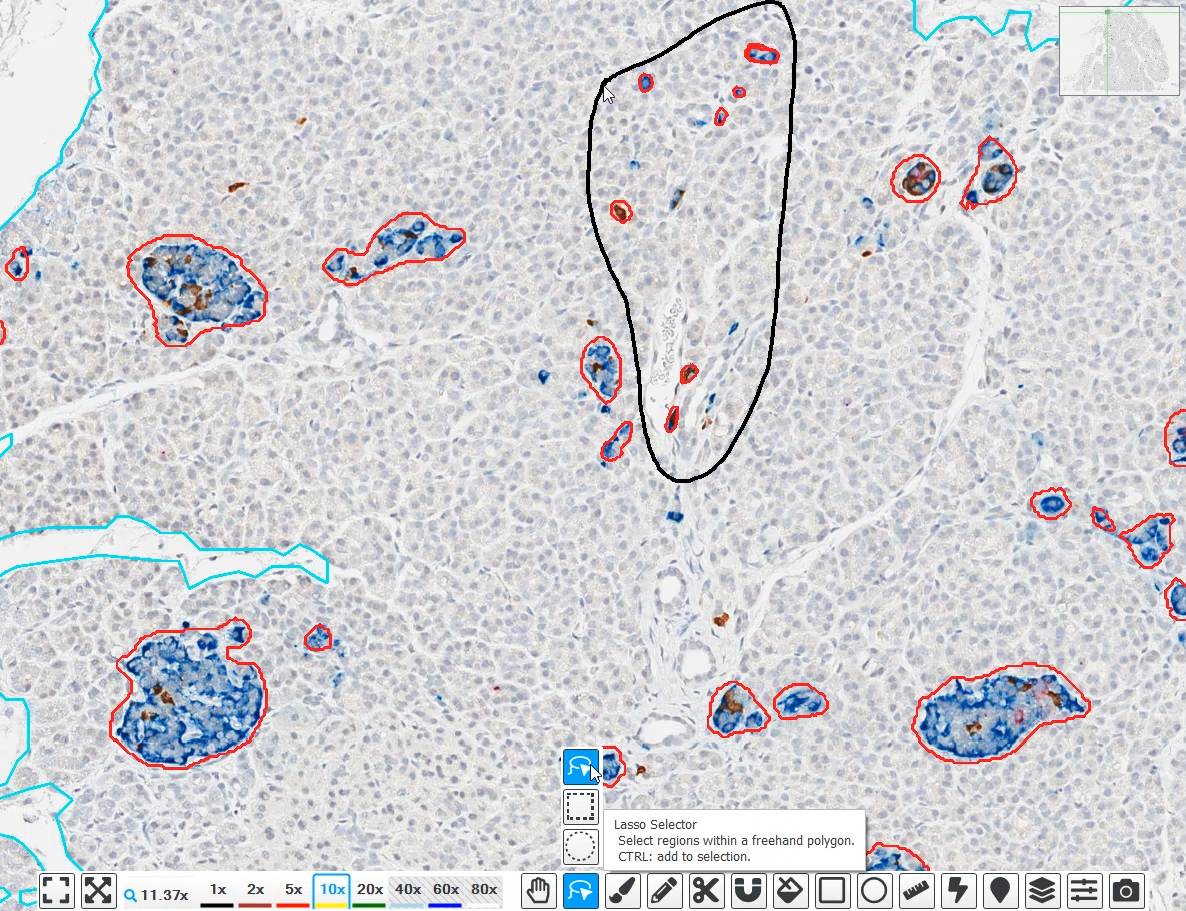
HALO®, HALO AI, and HALO Link 4.2 Features and Functionalities
In this blog post, you can learn about some of the new features in the 4.2 release, when to expect your chance to upgrade, and

Indica Labs’ Boston HALO® User Group Meeting 2026
12 May 2026 | Indica Labs is pleased to announce our Boston HALO® User Group Meeting at Le Méridien Boston Cambridge on 12 May from

Indica Labs’ Frankfurt HALO® User Group Meeting 2026
17 March 2026 | Indica Labs is pleased to announce our Frankfurt HALO® User Group Meeting at The Hilton Frankfurt Airport on 17 March from

Indica Labs’ London HALO® User Group Meeting 2025
9 December 2025 | Indica Labs is pleased to announce our London HALO® User Group Meeting at Hilton London Metropole on 9 December from 12:00

Deploy HALO® and HALO AI in a Fully Managed Cloud-Hosted Environment
Indica Labs offers optimized deployments of HALO and HALO AI software hosted in Amazon Web Services (AWS). We manage all aspects of implementation and ongoing maintenance to provide a highly performant, scalable, and secure AI-enabled HALO environment in the cloud. You maintain full ownership over your AWS account while our AWS Certified Solutions Architects manage your infrastructure and HALO software environments.
Learn more about benefits of cloud-hosted environments in our recent blog post. Or reach out today to our Cloud Services team by using the form below, to learn whether a managed cloud deployment is right for you.
Want to Learn More?
Fill out the form below to request a recorded or live demo of HALO AI or any of our other products or services.
You can also drop us an email at info@indicalab.com
Products & Services
Interested in purchasing or learning more about our products and services? Our highly trained application scientists are a couple of clicks away.
Software Maintenance & Support Coverage
Interested in purchasing an SMS plan? We would be happy to give you a quote.
Technical Support
Need technical support? Our IT specialists are here to help.