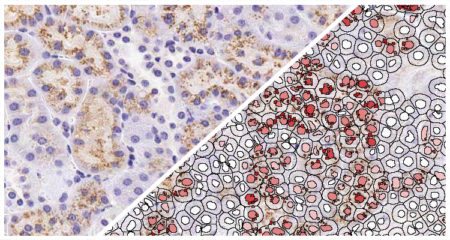
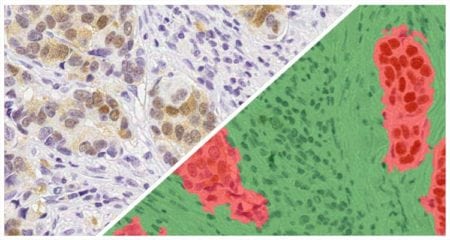
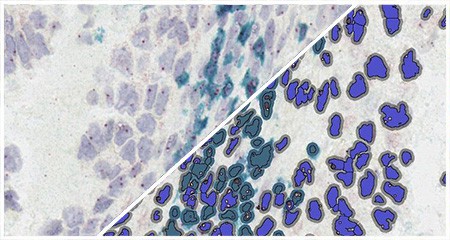
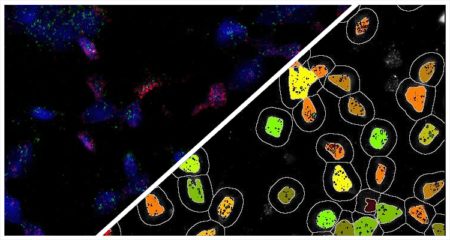
Indica Labs in situ hybridization (ISH) module is recognized as the fastest, most accurate, and user friendly analysis tool for quantifying nucleic acid probes across tissues and cell lines. This module measures chromogenic and/or silver-labelled DNA or RNA ISH probes on a cell-by-cell basis. This allows the user go rapidly contextualize the gene expression profile of every cell across a tissue. HALO ISH analysis is designed to work with up to three ISH probes and supports the H-score protocol for RNAscope as recommended by the manufacturer, ACD, a Bio-techne brand.
The module reports the total number of cells, total number of spots, total area of spots, the average number of spots per cell and per unit area of tissue, the average area of spots per cell and per unit area of tissue, as well as the number and percentage of cells with RNAscope score of 0, 1+, 2+, 3+ and 4+. When two probes are present, the module also reports the ratio of probe 1: probe 2. Histograms are automatically generated representing the frequency distribution of probes per cell [Figure 1] and RNAscope cell scoring [Figure 2] as shown below.

Figure 1 shows a distribution curve of probe number on a cell by cell basis, in this example most cells were shown to have between 0-5 copies per cell.
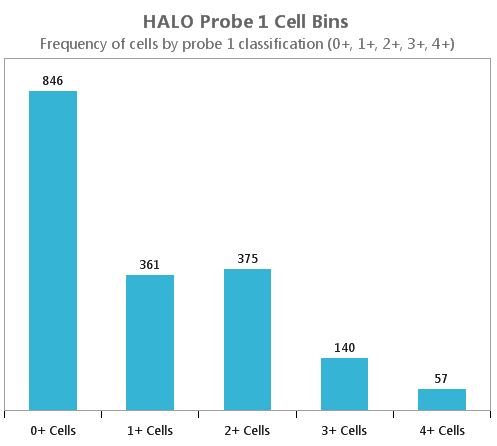 Figure 2. This bar chart shows the numbers of cells that were categorized as 0, 1+, 2+, 3+ and 4+ respectively. The chart shows that a slim majority of cells were positive for RNA probe expression.
Figure 2. This bar chart shows the numbers of cells that were categorized as 0, 1+, 2+, 3+ and 4+ respectively. The chart shows that a slim majority of cells were positive for RNA probe expression.
The ISH quantification module integrates seamlessly into the HALO® platform which is compatible with a number of third party systems and file formats.
File formats supported by the HALO image analysis platform:
- Non-proprietary (JPG, TIF, OME.TIFF)
- Nikon (ND2)
- 3D Histech (MRXS)
- Akoya (QPTIFF, component TIFF)
- Olympus / Evident (VSI)
- Hamamatsu (NDPI, NDPIS)
- Aperio (SVS, AFI)
- Zeiss (CZI)
- Leica (SCN, LIF)
- Ventana (BIF)
- Philips (iSyntax, i2Syntax)
- KFBIO (KFB, KFBF)
- DICOM (DCM*)
*whole-slide images

RNAscope™ Image Analysis Using HALO and HALO AI
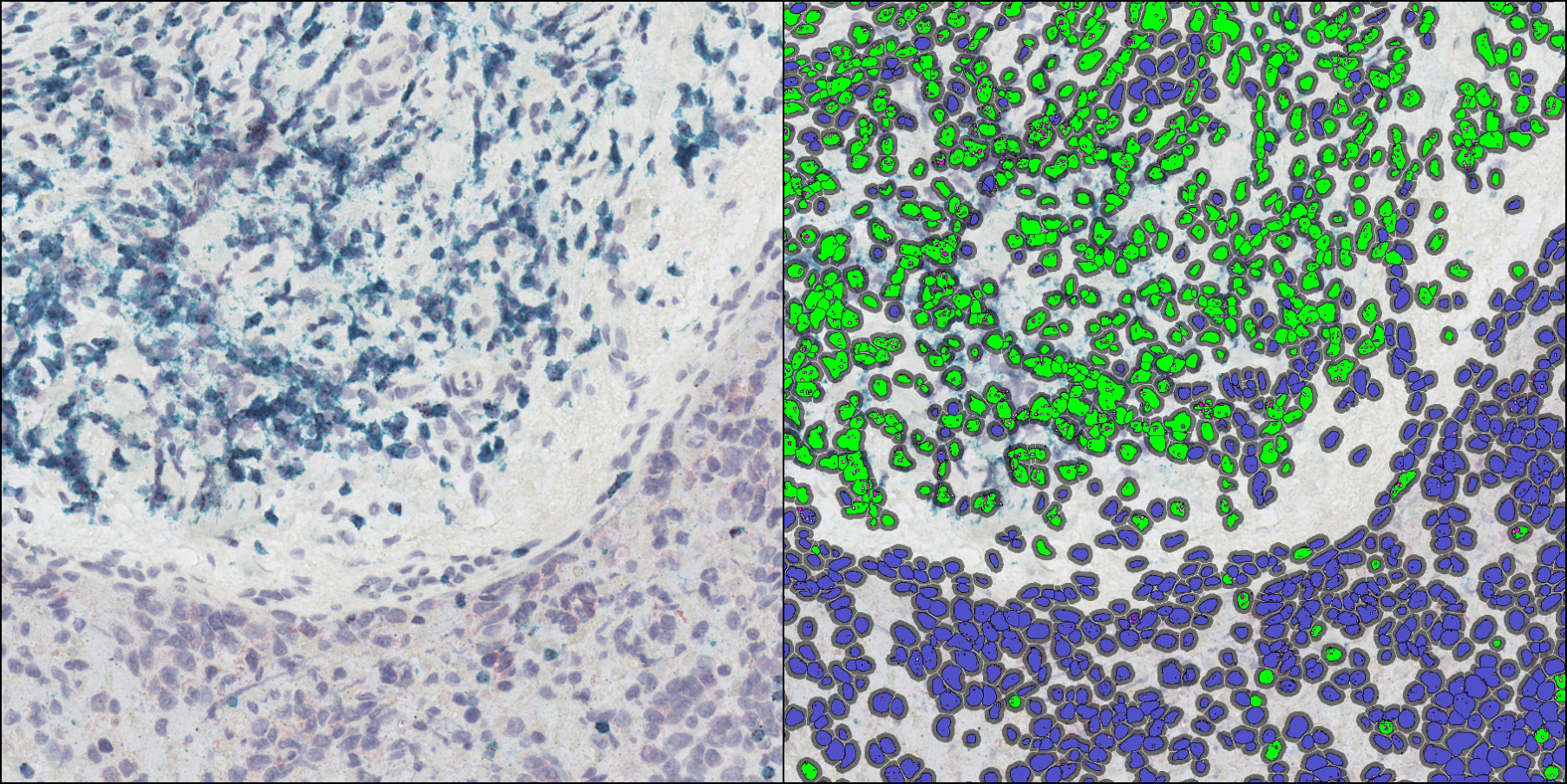
Using our ISH module for brightfield and our FISH module for fluorescence, we demonstrate how to quantify ISH signal from RNAscope™ assays based on ACD Bio’s recommended scoring guidelines.
Submit the form below to view the requested document

Glomeruli Quantification and Characterization Application Note
Don’t let varied stain selections hold back your digital image analysis of kidney sections! With HALO AI you can develop a tissue classifier that accurately detects and segments glomeruli in sections across a range of stains including H&E, trichrome, PAMS, and DAB with hematoxylin.
Submit the form below to view the requested document

RNAscope® Publications with the HALO® Image Analysis Platform
Read this publications list to see examples of research in areas from neuroscience to oncology using HALO to analyze RNAscope images.
Submit the form below to view the requested document

Quantitative RNAscope Image Analysis Guide
Advance your RNAscope image analysis by downloading the latest Quantitative RNAscope Image Analysis Guide from Indica Labs and ACD, a Bio-techne brand.
Submit the form below to view the requested document

Getting Started with RNAscope™ Image Analysis in HALO®
28 March 2023 | Join us for this 1-hour webinar to see a live demonstration of RNAscope image analysis using the HALO® platform from Indica
Masterclass Webinar: Optimizing RNAscope Image Analysis
05 May 2022 | In this 60-min webinar, Dr. Ghislaine Lioux will present solutions to common RNAscope image analysis challenges including how to optimize

Whole-slide Quantitative RNAscope Image Analysis: Applications and Methods
20 April 2022 | In this 2-hour event, learn from three experts in RNAscope image analysis with HALO® as they present their scientific research, share

HALO Image Analysis Masterclass Series: February-March 2021
Spring of 2021 | Indica Labs is excited to continue our HALO® Masterclass Webinar Series this winter. Each masterclass webinar will offer a deep dive
Publication Spotlight
The table below includes publications that cite the ISH Quantification module.
Your publication not on the list? Drop us an email to let us know about it!
| Title | Authors | Year | Journal | Application | HALO Modules | Product |
|---|---|---|---|---|---|---|
| EBV+†tumors exploit tumor cell-intrinsic and -extrinsic mechanisms to produce regulatory T cell-recruiting chemokines CCL17 and CCL22 | Jorapur A, Marshall L, Jacobson S, Xu M, Marubayashi S, Zibinsky M, Hu D, Robles O, Jackson J, Baloche V, Busson P, Wustrow D, Brockstedt D, Talay O, Kassner P, Cutler G | 2022 | PLOS Pathogens | Oncology | ISH/FISH | HALO |
| AAV9/MFSD8 gene therapy is effective in preclinical models of neuronal ceroid lipofuscinosis type 7 disease | Chen X, Dong T, Hu Y, Shaffo F, Belur N, Mazzulli J, Gray S | 2022 | The Journal of Clinical Investigation | Neuroscience | Area Quantification, ISH/FISH | HALO |
| Genome Sequence and Experimental Infection of Calves with Bovine Gammaherpesvirus 4 (BoHV-4) | Bauermann F, Falkenberg S, Martins M, Dassanayake R, Neill J, Ridpath J, Silveira S, Palmer M, Buysse A, Mohr A, Flores E, Diel D | 2022 | Research Square | Infectious Disease, Other | ISH/FISH | HALO |
| Vertical transmission of attaching and invasive E. coli from the dam to neonatal mice predisposes to more severe colitis following exposure to a colitic insult later in life | Brand M, Proctor A, Hostetter J, Zhou N, Friedberg I, Jergens A, Phillips G, Wannemuehler M | 2022 | PLOS ONE | Other | ISH/FISH | HALO |
| Epicardium-derived cells organize through tight junctions to replenish cardiac muscle in salamanders | Eroglu E, Yen C, Witman N, Elewa A, Araus A, Wang H, Szattler T, Umeano C, Sohlmer J, Goedel A, Simon A, Chien K | 2022 | Nature Cell Biology | Myology | ISH/FISH, ISH-IHC/FISH-IF | HALO |
| TGF?1 Induces Senescence and Attenuated VEGF Production in Retinal Pericytes | Avarmovic D, Archaimbault S, Kemble A, Gruener S, Lazendic M, Westenskow P | 2022 | Biomedicines | Other | ISH/FISH | HALO |
| Satellite repeat RNA expression in epithelial ovarian cancer associates with a tumor immunosuppressive phenotype | Porter R, Sun S, Flores M, Berzolla E, You E, Phillips I, KC N, Desai N, Tai E, Szabolcs A, Lang E, Pankaj A, Raabe M, Thapar V, Xu K, Nieman L, Rabe D, Kolin D, Stover E, Pepin D, Stott S, Deshpande V, Liu J, Solovyov A, Matulonis U, Greenbaum B, Ting D | 2022 | The Journal of Clinical Investigation | Immuno-oncology | ISH/FISH, ISH-IHC/FISH-IF | HALO |
| Distinct Encoding of Reward and Aversion by Peptidergic BNST Inputs to the VTA | Soden M, Yee J, Cuevas B, Rastani A, Elum J, Zweifel L | 2022 | Frontiers in Neural Circuits | Neuroscience | ISH/FISH | HALO |
| Hyperpolarised†13C-MRI identifies the emergence of a glycolytic cell population within intermediate-risk human prostate cancer | Sushentsev N, McLean M, Warren A, Benjamin A, Brodie C, Frary A, Gill A, Jones J, Kaggie J, Lamb B, Locke M, Miller J, Mills I, Priest A, Robb F, Shah N, Schulte R, Graves M, Gnanapragasam V, Brindle K, Barrett T, Gallacher F | 2022 | Nature Communications | Oncology | ISH/FISH, Membrane, Multiplex IHC | HALO |
| Adult re-expression of IRSp53 rescues NMDA receptor function and social behavior in IRSp53-mutant mice | Noh Y, Yook C, Kang J, Lee S, Kim Y, Yang E, Kim H, Kim E | 2022 | Communications Biology | Neuroscience | ISH/FISH, Figure Maker | HALO |
| The NALCN channel regulates metastasis and nonmalignant cell dissemination | Rahrmann E, Shorthouse D, Jassim A, Hu L, Ortiz M, Mahler-Araujo B, Vogel P, Paez-Ribes M, Fatemi A, Hannon G, Iyer R, Blundon J, Lourenco F, Kay J, Nazarian R, Hall B, Zakharenko S, Winton D, Zhu L, Gilbertson R | 2022 | Nature Genetics | Oncology | ISH/FISH, Multiplex IHC | HALO |
| Relaxin/insulin-like family peptide receptor 4 (Rxfp4) expressing hypothalamic neurons Q8 modulate food intake and preference in mice | Lewis J, Woodward O, Nuzzaci D, Smith C, Adriaenssens A, Billing L, Brighton C, Phillips B, Tadross J, Kinston S, Ciabatti E, Gottgens B, Tripodi M, Mornigold D, Baker D, Gribble F, Reimann F | 2022 | Molecular Metabolism | Metabolism | ISH/FISH, Highplex FL | HALO |
| Regional epithelial cell diversity in the small intestine of pigs | Wiarda J, Becker S, Sivasankaran S, Loving C, | 2022 | Journal of Animal Science | Gastroenterology | ISH/FISH, Figure Maker | HALO |
| Single cell analysis of cribriform prostate cancer reveals cell intrinsic and tumor microenvironmental pathways of aggressive disease | Wong H, Sheng Q, Hesterberg A, Croessmann S, Rios B, Giri K, Jackson J, Miranda A, Watkins E, Schaffer K, Donahue M, Winkler E, Penson D, Smith J, Herrell S, Luckenbaugh A, Barocas D, Kim Y, Graves D, Giannico G, Rathmell J, Park B, Gordetsky J, Hurley P | 2022 | Nature Communications | Oncology | ISH/FISH | HALO |
| Systemic Nos2 Depletion and Cox inhibition limits TNBC disease progression and alters lymphoid cell spatial orientation and density | Somasundaram V, Ridnour L, Cheng R, Walke A, Kedei N, Bhattacharyya D, Wink A, Edmondson E, Butcher D, Warner A, Dorsey T, Scheiblin D, Heinz W, Bryant R, Kinders R, Lipkowitz S, Wong S, Pore M, Hewitt S, McVicar D, Anderson S, Chang J, Glynn S, Ambs S, Lockett S, Wink D | 2022 | Redox Biology | Oncology | Classifer, ISH/FISH, Spatial Analysis, Registration, ISH-IHC/FISH-IF | HALO |
| Glutaminase 2 Knockdown Reduces Hyperammonemia and Associated Lethality of Urea Cycle Disorder Mouse Model | Mao X, Chen H, Lin A, Kim S, Burczynski M, Na E, Halasz G, Sleeman M, Murphy A, Okamoto H, Cheng X | 2022 | Journal of Inherited Metabolic Disease | Metabolism | Area Quantification, ISH/FISH | HALO |
| What Happens to the Preserved Renal Parenchyma After Clamped Partial Nephrectomy? | Xiong L, Nguyen J, Peng Y, Zhou Z, Ning K, Jia N, Nie J, Wen D, Wu Z, Roversi G, Palacios D, Abramczyk E, Munoz-Lopez C, Campbell J, Cao Y, Li Wencai, Zhang X, He Z, Li X, Huang J, Shou J, Wu J, Chen M, Chen X, Zheng J, Xu C, Zhong W, Li Z, Dong W, Zhou J, Zhang H, Luo J, Liu J, Sun G, Han H, Guo S, Dong P, Zhou F, Yu C, Campbell S, Zhang Z | 2022 | European Urology | Other | Area Quantification, ISH/FISH | HALO |
| Adolescent intermittent ethanol exposure produces Sex-Specific changes in BBB Permeability: A potential role for VEGFA | Vore A, Barney T, Deak M, Varlinskaya E, Deak T | 2022 | Brain, Behavior, and Immunity | Neuroscience | ISH/FISH | HALO |
| Relationship between impaired BMP signalling and clinical risk factors at early-stage vascular injury in the preterm infant | Heydarian M, Oak P, Zhang X, Kamgari N, Kindt A, Koschlig M, Pritzke T, Gonzalez-Rodriguez E, Forster K, Morty R, Hafner F, Hubener C, Flemmer A, Yildirim A, Sudheendra D, Tian X, Petrera A, Kirsten H, Ahnert P, Morrell N, Desai T, Sucre J, Spiekerkoetter E, Hilgendorff A | 2022 | Thorax | Other | ISH/FISH | HALO |
| Spatiotemporal expression pattern of Progesterone Receptor Component (PGRMC) 1 in endometrium from patients with or without endometriosis or adenomyosis | Thieffry C, Van Wynendaele M, Samain L, Tyteca D, Pierreux C, Marbaix E, Henriet P | 2022 | The Journal of Steroid Biochemistry and Molecular Biology | Other | ISH/FISH, Highplex FL | HALO, HALO AI |
| Quantitative Imaging Analysis of the Spatial Relationship between Antiretrovirals, Reverse Transcriptase Simian-Human Immunodeficiency Virus RNA, and Fibrosis in the Spleens of Nonhuman Primates | Devanathan A, White N, Desyaterik Y, De la Cruz G, Nekorchuk M, Terry M, Busman-Sahay K, Adamson L, Luciw P, Fedoriw Y, Estes J, Rosen E, Kashuba A | 2022 | Antimicrobial Agents and Chemotherapy | Infectious Disease | ISH/FISH | HALO |
| CD123-directed allogeneic chimeric-antigen receptor T-cell therapy (CAR-T) in blastic plasmacytoid dendritic cell neoplasm (BPDCN): Clinicopathological insights | Pemmaraju N, Wilson N, Senapati J, Economides M, Guzman M, Neelapu S, Kazemimood R, Davis R, Jain N, Khoury J, Sugita M, Cai T, Smith J, Frattini M, Garton A, Roboz G, Konopleva M | 2022 | Leukemia Research | Immuno-oncology | ISH/FISH | HALO |
| Localization of macrophage subtypes and neutrophils in the prostate tumor microenvironment and their association with prostate cancer racial disparities | Maynard J, Godwin T, Lu J, Vidal I, Lotan T, De Marzo A, Joshu C, Sfanos K | 2022 | The Prostate | Oncology | ISH/FISH, Multiplex IHC | HALO |
| Molecular identification of spatially distinct anabolic responses to mechanical loading in murine cortical bone | Chlebek C, Moore J, Ross F, van der Meulen M | 2022 | Journal of Bone and Mineral Research | Other | ISH/FISH | HALO |
| Sex-specific effects of ethanol consumption in older Fischer 344 rats on microglial dynamics and A_(1-42) accumulation | Marsland P, Vore A, DaPrano E, Paluch J, Blackwell A, Varlinskaya E, Deak T | 2022 | Alcohol | Neuroscience | ISH/FISH | HALO |
| Estrogen receptor beta activity contributes to both tumor necrosis factor alpha expression in the hypothalamic paraventricular nucleus and the resistance to hypertension following angiotensin II in female mice | Woods C, Contoreggi N, Johnson M, Milner T, Wang G, Glass M, | 2022 | Neurochemistry International | Neuroscience | ISH/FISH | HALO |
| Angiotensin II Infusion Results in Both Hypertension and Increased AMPA GluA1 Signaling in Hypothalamic Paraventricular Nucleus of Male but not Female Mice | Wang G, Woods C, Johnson M, Milner T, Glass M | 2022 | Neuroscience | Neuroscience | ISH/FISH | HALO |
| Multiregional single-cell dissection of tumor and immune cells reveals stable lock-and-key features in liver cancer | Ma L, Heinrich S, Wang L, Keggenhoff F, Khatib S, Forgues M, Kelly M, Hewitt S, Saif A, Hernandez J, Mabry D, Kloeckner R, Greten T, Chaisaingmongkol J, Ruchirawat M, Marquardt J, Wang X | 2022 | Nature Communications | Oncology | ISH/FISH | HALO |
| Autologous T cell therapy for MAGE-A4+ solid cancers in HLA-A*02+ patients: a phase 1 trial | Hong D, Van Tine B, Biswas S, McAlpine C, Johnson M, Olszanski A, Clarke J, Araujo D, Blumenschein G, Kebriaei P, Lin Q, Tipping A, Sanderson J, Wang R, Trivedi T, Annareddy T, Bai J, Rafail S, Sun A, Fernandes L, Navenot J-M, Bushman F, Everett J, Karadeniz D, Broad R, Isabelle M, Naidoo R, Bath N, Betts G, Wolchinsky Z, Batrakou D, Van Winkle E, Elefant E, Ghobadi A, Cashen A, Grand'Maison A, McCarthy P, Fracasso P, Norry E, Williams D, Druta M, Liebner D, Odunsi K, Butler M | 2023 | Nature Medicine | Immuno-oncology | ISH/FISH, Spatial Analysis, Highplex FL | HALO |
| Combined KRASG12C and SOS1 inhibition enhances and extends the anti-tumor response in KRASG12C-driven cancers by addressing intrinsic and acquired resistance | Thatikonda V, Lu H, Jurado S, Kostyrko K, Bristow C, Bosch K, Feng N, Gao S, Gerlach D, Gmachi M, Lieb S, Jeschko A, Machado A, Marszalek E, Mahendra M, Jaeger P, Sorokin A, Strauss S, Trapani F, Kopetz S, Vellano C, Petronczki M, Kraut N, Heffernan T, Marszalek J, Pearson M, Waizenegger I, Hofmann M | 2023 | bioRxiv | Oncology | ISH/FISH, Multiplex IHC, Highplex FL, Figure Maker | HALO |
| Prefrontal cortical protease TACE/ADAM17 is involved in neuroinflammation and stress-related eating alterations | Sharafeddin F, Ghaly M, Simon T, Ontiveros-Angel P, Figueroa J | 2023 | bioRxiv | Neuroscience | ISH/FISH | HALO |
| Long noncoding RNA LINC01594 inhibits the CELF6-mediated splicing of oncogenic CD44 variants to promote colorectal cancer metastasis. | Liu B, Song A, Gui P, Wang J, Pan Y, Li C, Li S, Zhang Y, Jiang T, Xu Y, Huo F, Pei D, Song J | 2023 | Research Square | Oncology | ISH/FISH | HALO |
| CD47-Dependent Regulation of Immune Checkpoint Gene Expression and MYCN mRNA Splicing in Murine CD8 and Jurkat T Cells | Kaur S, Awad D, Finney R, Meyer T, Singh S, Cam M, Karim B, Warner A, Roberts D | 2023 | International Journal of Molecular Sciences | Immuno-oncology | ISH/FISH | HALO |
| SARS-CoV-2 infection induces DNA damage, through CHK1 degradation and impaired 53BP1 recruitment, and cellular senescence | Gioia U, Tavella S, Martinez-Orellana P, Cicio G, Colliva A, Ceccon M, Cabrini M, Henriques A, Fumagalli V, Paldino A, Presot E, Rajasekharan S, Iacomino N, Pisati F, Matti V, Sepe S, Conte M, Barozzi S, Lavagnino Z, Carletti T, Volpe M, Cavalcante P, Iannacone M, Rampazzo C, Bussani R, Tripodo C, Zacchigna S, Marcello A, di Fagagna F | 2023 | Nature Cell Biology | Infectious Disease | ISH/FISH, Highplex FL | HALO |
| A cellular and molecular spatial atlas of dystrophic muscle | Stec M, Su Q, Adler C, Zhang L, Golann D, Khan N, Panagis L, Villalta S, Ni M, Wei Y, Walls J, Murphy A, Yancopoulos G, Atwal G, Kleiner S, Halasz G, Sleeman M | 2023 | Proceedings of the National Academy of Sciences | Myology | ISH/FISH, Highplex FL | HALO |
Related HALO Modules
Separate multiple tissue classes across a tissue using a learn-by-example approach. Can be used in conjunction with all other modules (fluorescent and brightfield) to select specific tissue classes for further analysis.
Learn MoreQuantify IHC positivity and spot number, optical density and co-expression of one or two brightfield RNA or DNA probes on a per cell basis. Includes support for brightfield RNAscope assays.
Learn MoreQuantify number, intensity and co-expression of an unlimited number of fluorescently labeled RNA or DNA probes on a per cell basis. Includes support for fluorescent RNAscope assays.
Learn MoreUse the arrows above to view additional related modules
Want to Learn More?
Fill out the form below to request information about any of our software products.
You can also drop us an email at info@indicalab.com