Particularly useful to those involved in the immune-oncology field, the Spatial Analysis module offers a suite of subsequent analysis tools which can be used to identify proximity and relative spatial distribution of objects, cells, and/or features across single tissues or serial sections. This module is embedded within the HALO platform and can be used in conjunction with any of our cell-based analysis modules for brightfield or fluorescence.
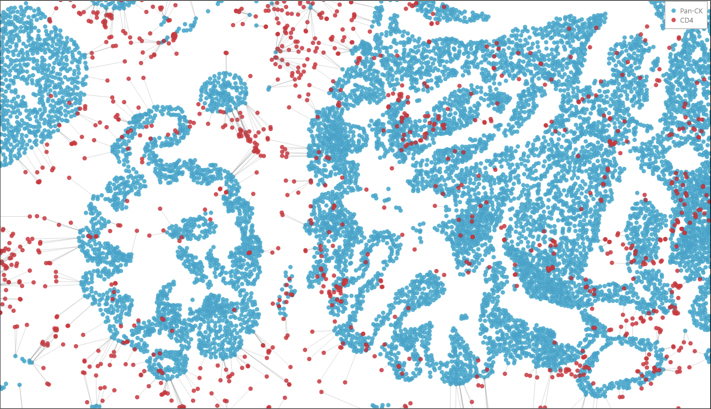

Nearest Neighbor
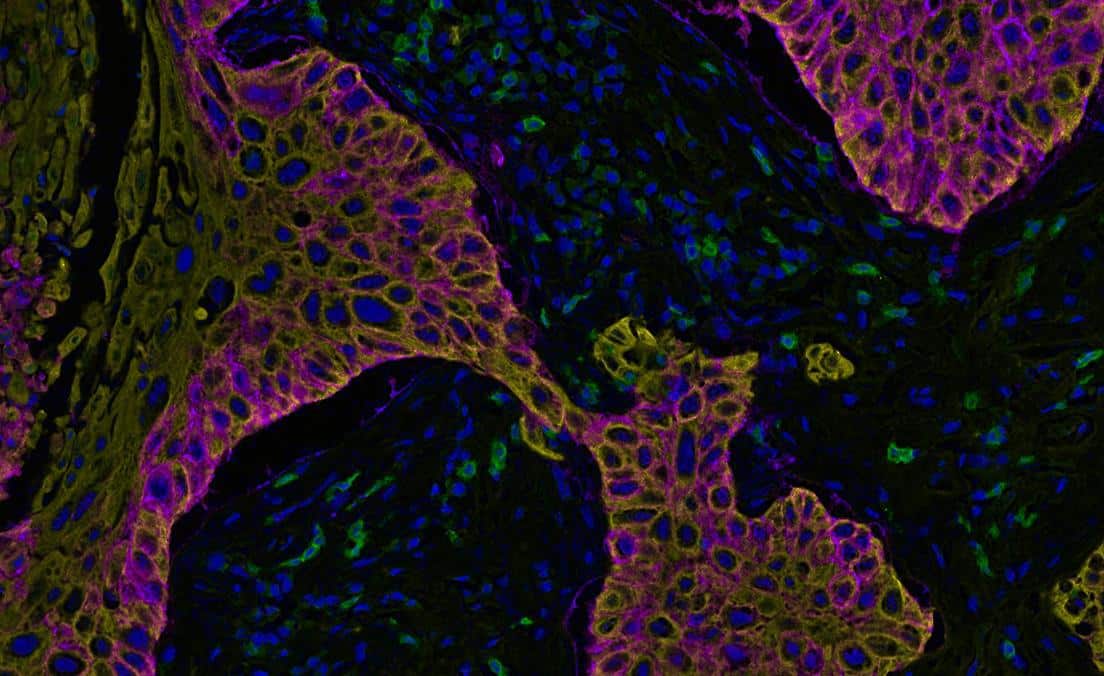
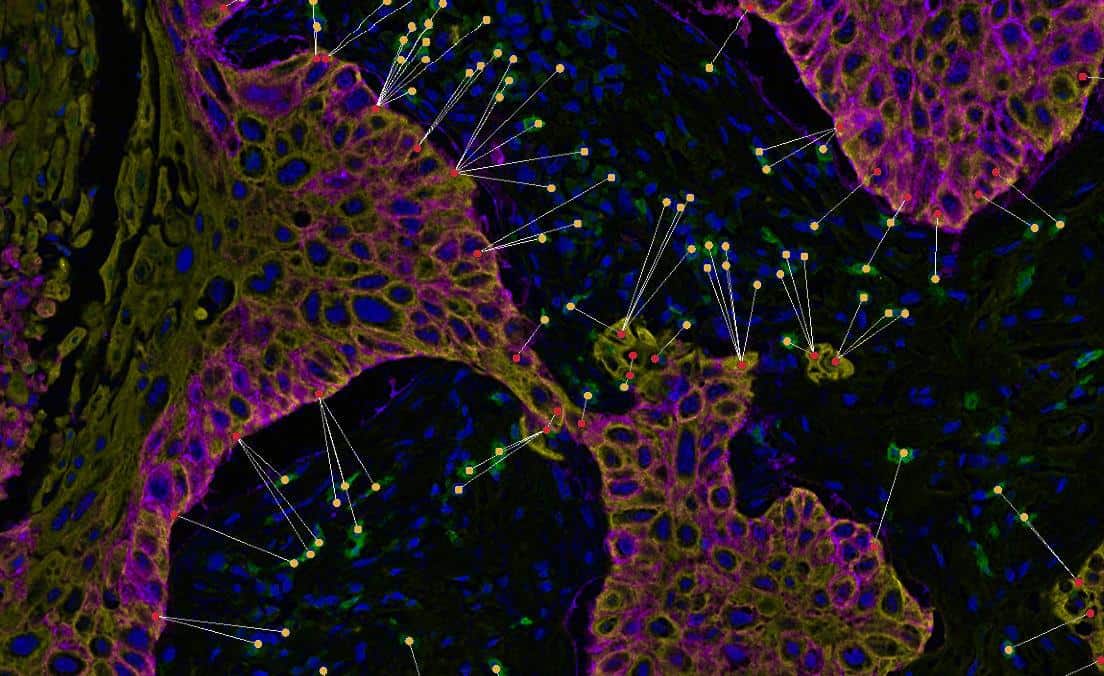
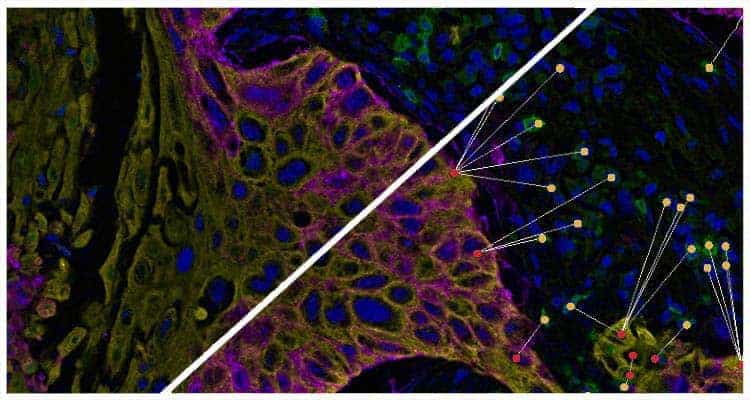
Determine the average distance and number of unique neighbors between any two cell or object populations. The example below depicts pan cytokeratin positive cells (blue) and CD4 positive cells (red). Grey lines connect each CD4 positive cell to the nearest pan cytokeratin positive cell.
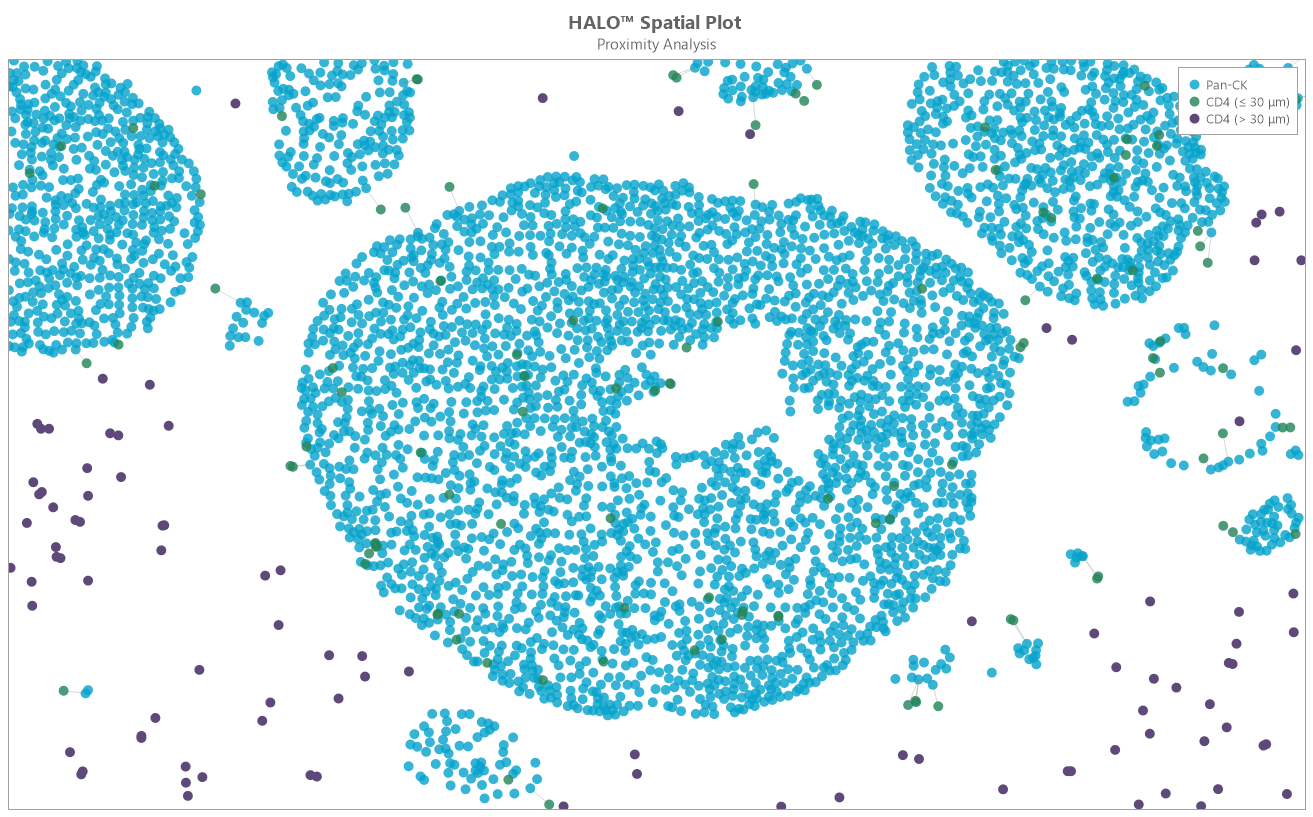
Proximity Analysis
Calculate the number of objects or cells within a certain distance of another object or cell. In the example below, the distance is set to 30 microns. CD8 positive cells within 30 microns of a pan cytokeratin positive cell (blue) are labeled green and CD8 positive cells greater than 30 microns from pan cytokeratin positive cell are labeled purple. A corresponding proximity histogram is automatically generated.
Infiltration Analysis
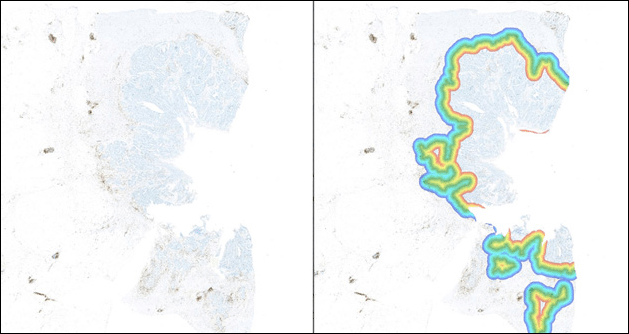
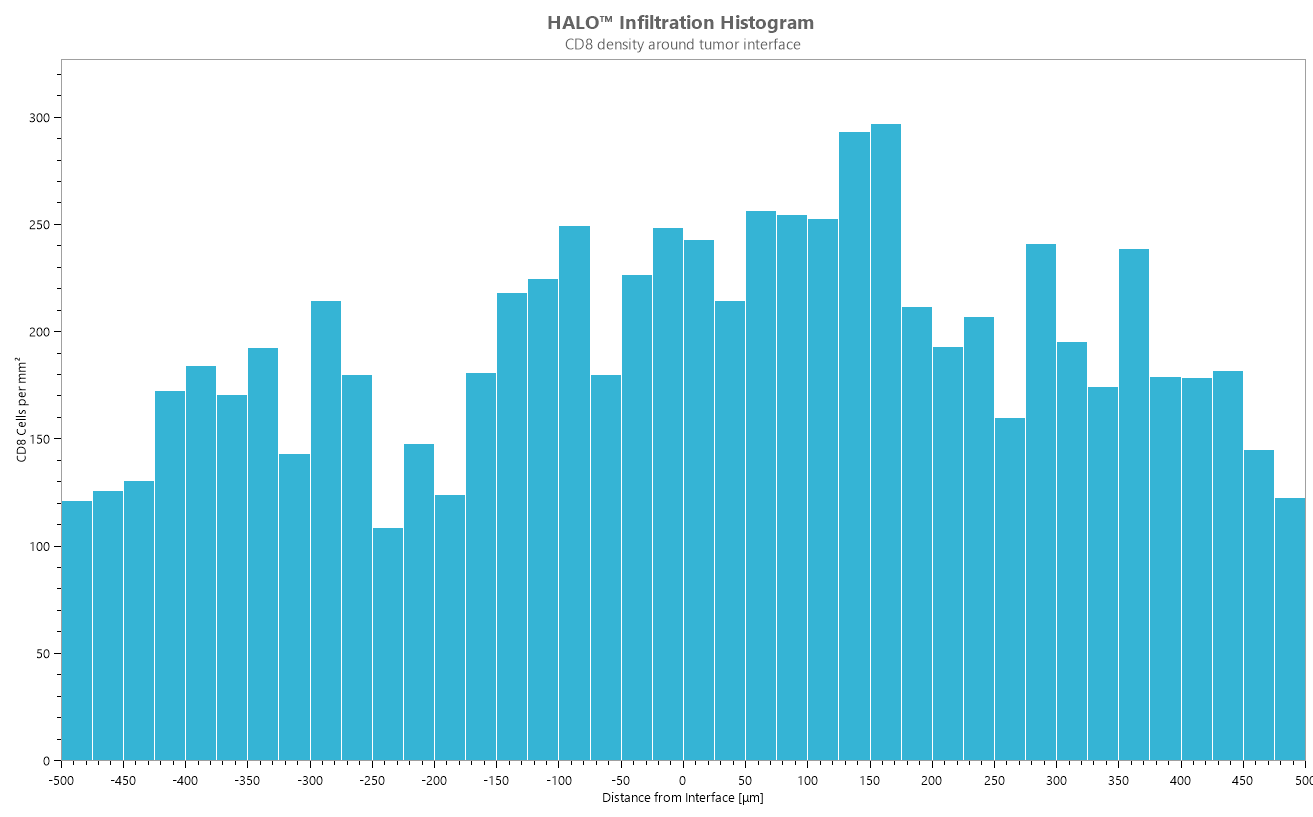
Determine the number of objects or cells within a set range of an annotated region of interest. Depicted below is a whole slide image stained with CD8. The tumor boundary has been defined/annotated in green. The infiltration analysis tool defines the invasive margin inside of tumor (yellow and red) and the invasive margin outside of the tumor (blue and purple) automatically. In this example, the distance is set to 500 microns around the annotated tumor boundary. CD8 positive cells are quantified within the margin. A histogram is automatically generated to reflect CD8+ cell density inside the tumor boundary (-1 to -500), at the tumor boundary (0), and outside the tumor boundary (+1 to +500) as shown in the image below.
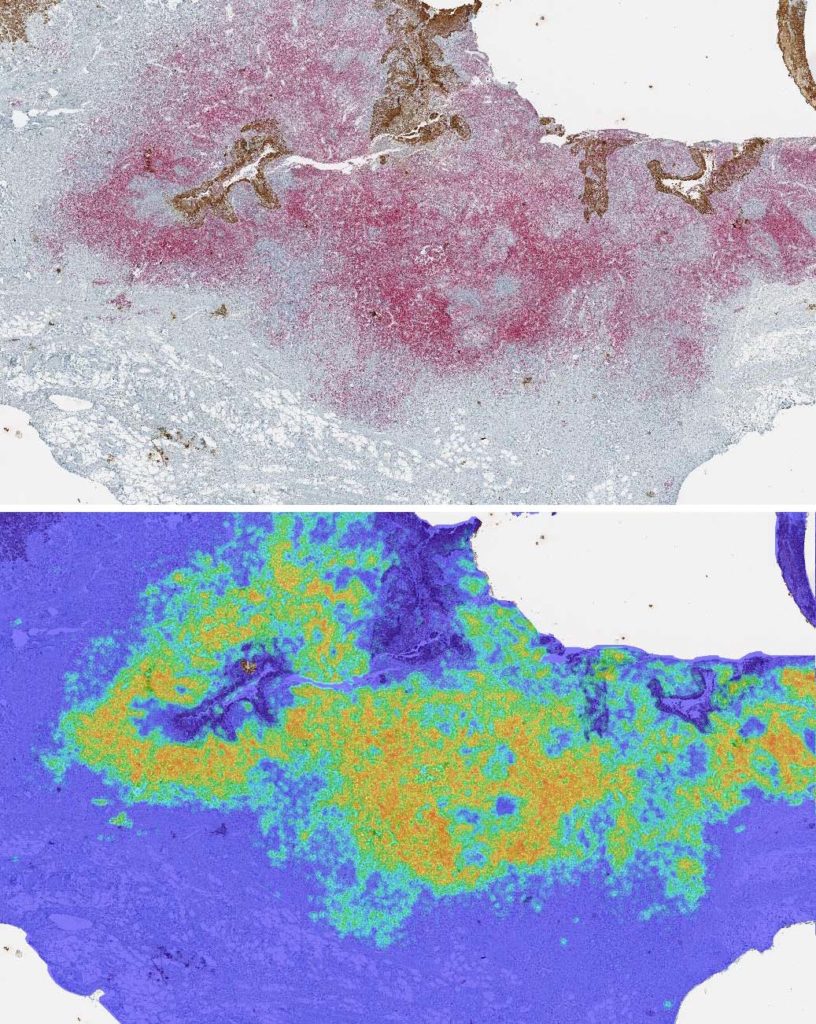

Density Heatmaps
The density heatmap spatial analysis algorithm measures the density of a selected cell population from your object data within a certain radius. The scaling and colors of the markup are customizable. Reported in the summary data, the entire tissue area is analyzed and all cells of the desired population are counted. The average cell population and the average cell population density is calculated within the user-defined radius, as well as the minimum and maximum densities.
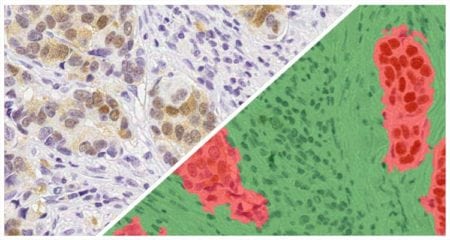
The tissue below is stained with an immune cell marker (red stain) and a tumor marker (DAB stain). The image was analyzed with the Multiplex IHC module to detect positivity and produce object data. The density heatmap analysis was used with a radius of 25 microns, measuring the density of immune cell positivity across the whole slide. The markup image, with adjustable transparency, depicts areas of higher density in red and orange, moderate density in yellow, and areas of lower density in green and blue.
File formats supported by the HALO image analysis platform:
- Non-proprietary (JPG, TIF, OME.TIFF)
- Nikon (ND2)
- 3D Histech (MRXS)
- Akoya (QPTIFF, component TIFF)
- Olympus / Evident (VSI)
- Hamamatsu (NDPI, NDPIS)
- Aperio (SVS, AFI)
- Zeiss (CZI)
- Leica (SCN, LIF)
- Ventana (BIF)
- Philips (iSyntax, i2Syntax)
- KFBIO (KFB, KFBF)
- DICOM (DCM*)
*whole-slide images

Fusion of Spectrally Unmixed Image Tiles into Whole Slide TIFs and HALO Image Analysis
In this application note, we describe how HALO can convert spectrally unmixed image tiles into whole slide images for analysis of the tumor microenvironment.
Submit the form below to view the requested document

Characterizing the TME using IMC and HALO® Image Analysis
See in this app note how automated analysis of highly multiplexed IMC images using HALO and HALO AI yields rich cellular and spatial data from a streamlined workflow.
Submit the form below to view the requested document

Spatial Analysis Application Note
In this application note, we describe two HALO analysis tools that are used to quantify the distribution of immune cells relative to tumor cells in the tumor microenvironment, Proximity Analysis and Tumor Infiltration Analysis
Submit the form below to view the requested document

HiPlex RNAscope™ Application Note
Using our FISH module we demonstrate how to perform 12-plex RNAscope image analysis using ACD’s HiPlexv2 assay.
Submit the form below to view the requested document

RNAscope™ Image Analysis Using HALO and HALO AI
Using our ISH module for brightfield and our FISH module for fluorescence, we demonstrate how to quantify ISH signal from RNAscope™ assays based on ACD Bio’s recommended scoring guidelines.
Submit the form below to view the requested document
Depicting TME Cellular Architecture Using Hyperplex IF and Automated Image Analysis
Download our eBook in collaboration with Lunaphore to learn more about how Pharma Services depicted TME cellular architecture using hyperplex IF and automated image analysis.
Submit the form below to view the requested document
Lunaphore COMET Hyperplex IF and HALO® Image Analysis of the TME
Download our collaborative poster with Lunaphore to see how HALO and HALO AI, alongside the Lunaphore COMET instrument, enable a high throughput workflow for phenotypic and spatial analysis of cells in the tumor microenvironment.
Submit the form below to view the requested document
Characterizing the TME using IMC and HALO® Image Analysis
See in this app note how automated analysis of highly multiplexed IMC images using HALO and HALO AI yields rich cellular and spatial data from a streamlined workflow.
Submit the form below to view the requested document
Automated Tumor Budding Quantification in Colorectal Carcinoma H&E Images
Learn about quantification of tumor budding in colorectal carcinoma using HALO and HALO AI.
Submit the form below to view the requested document
Quantitative Image Analysis of a Combined FISH-IF Assay
Learn how to detect protein and RNA in a single assay using ACD’s Co-Detection workflow and HALO image analysis.
Submit the form below to view the requested document

Spatial Characterization of the TME with Cell DIVE Multiplexed Imaging and HALO Image Analysis
Check out our collaborative poster with Cell Signaling Technology and Leica Microsystems to learn about a combined workflow for spatial analysis of the TME.
Submit the form below to view the requested document
Spatial Multiplex Profiling of Immune Markers with the RNAscope Hiplex v2 Assay
Download this poster to learn about combining ACD’s HiPlex v2 in situ hybridization assay with HALO image analysis.
Submit the form below to view the requested document

Quantitative RNAscope Image Analysis Guide
Advance your RNAscope image analysis by downloading the latest Quantitative RNAscope Image Analysis Guide from Indica Labs and ACD, a Bio-techne brand.
Submit the form below to view the requested document

A role for HALO® in characterizing cell heterogeneity across organ systems: from the liver to the brain
12 October 2023 | Please join us for this 1-hour webinar to learn about characterizing cell heterogeneity with HALO to gain new biological insights.
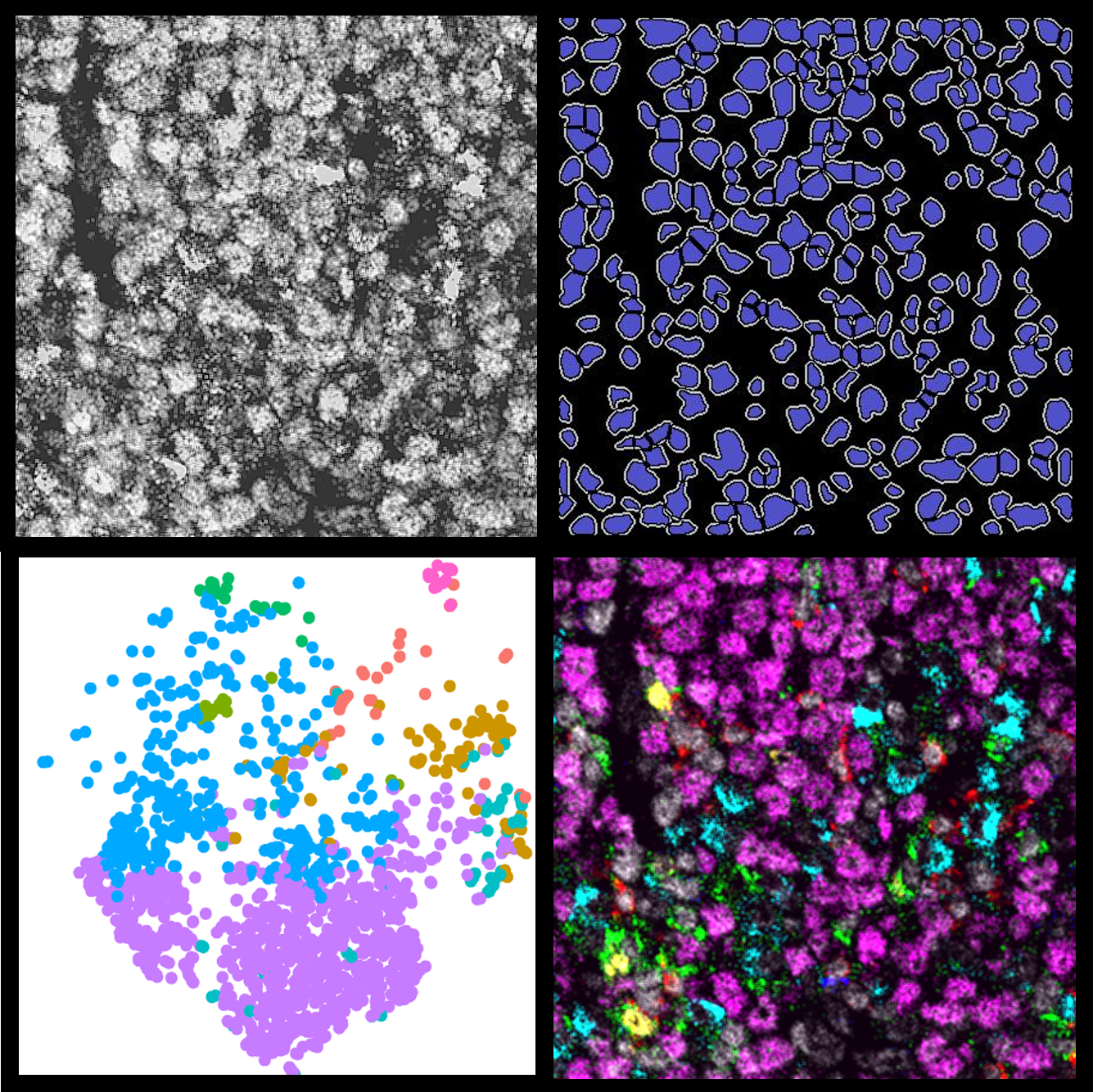
Advanced Multiplex and Spatial Analysis Methods using HALO Data
9 June 2022 | In this 90-minute webinar, Learn about an open-source MIBI image analysis pipeline using HALO data and how to validate a multiplex

Whole-slide Quantitative RNAscope Image Analysis: Applications and Methods
20 April 2022 | In this 2-hour event, learn from three experts in RNAscope image analysis with HALO® as they present their scientific research, share
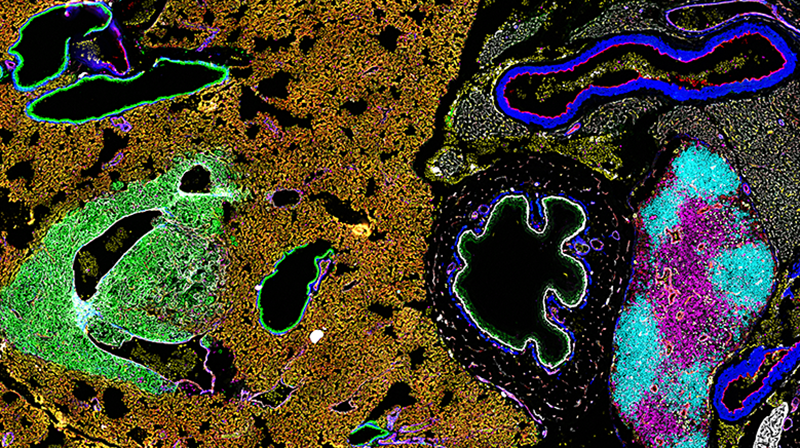
Optimizing quantitative analysis of highly multiplexed CODEX images in HALO
31 March 2022 | In this webinar, Dr. Noemi Kedei, a staff scientist at the Collaborative Protein Technology Resource core of the Center
Publication Spotlight
The table below includes publications that cite the Spatial Analysis module.
Your publication not on the list? Drop us an email to let us know about it!
| Title | Authors | Year | Journal | Application | HALO Modules | Product |
|---|---|---|---|---|---|---|
| Progranulin mediates immune evasion of pancreatic ductal adenocarcinoma through regulation of MHCI expression | Cheung P, Yang J, Fang R, Borgers A, Krengel K, Stoffel A, Althoff K, Yip C, Siu E, Ng L, Lang K, Cham L, Engel D, Soun C, Cima I, Scheffler B, Striefler J, Sinn M, Bahra M, Pelzer U, Oettle H, Markus P, Smeets E, Aarntzen E, Savvatakis K, Liffers S, Lueong S, Neander C, Bazarna A, Zhang X, Paschen A, Crawford H, Chan A, Cheung S, Siveke J | 2022 | Nature Communications | Oncology | Cytonuclear, Spatial Analysis, Highplex FL | HALO |
| Tumor cell-derived exosomes deliver TIE2 protein to macrophages to promote angiogenesis in cervical cancer | Du S, Qian J, Tan S, Li W, Liu P, Zhao J, Zeng Y, Xu L, Wang Z, Cai J | 2022 | Cancer Letters | Oncology | Area Quantification, Cytonuclear, Spatial Analysis, Highplex FL | HALO |
| Intratumoural spatial distribution of S100B?+?folliculostellate cells is associated with proliferation and expression of FSH and ER? in gonadotroph tumours | Ilie M, Vasiljevic A, Chanal M, Gadot N, Chinezu L, Jouanneau E, Hennino A, Raverot G, Bertolino P | 2022 | Acta Neuropathologica Communications | Oncology | Cytonuclear, Spatial Analysis | HALO |
| Interspatial Distribution of Tumor and Immune Cells in Correlation with PD-L1 in Molecular Subtypes of Gastric Cancers | Dislich B, Mertz K, Gloor B, Langer R | 2022 | Cancers | Immuno-oncology | Multiplex IHC, Spatial Analysis, Figure Maker | HALO |
| Peritumoral TIGIT+CD20+†B cell infiltration indicates poor prognosis but favorable adjuvant chemotherapeutic response in gastric cancer | Liu H, Wu J, Xu X, Wang H, Zhang C, Yin S, He Y | 2022 | International Immunopharmacology | Oncology | Cytonuclear, Spatial Analysis, Highplex FL | HALO |
| SPOP promotes cervical cancer progress by inducing PD-1 move away from PD-L1 in spatial localization | Wu J, Wu Y, Guo Q, Chen S, Wang S, Zhu J, Wu X, Ju X | 2022 | Research Square | Oncology | Cytonuclear, Spatial Analysis, Highplex FL, TMA | HALO |
| Higher proportions of CD39+ tumorresident cytotoxic T cells predict recurrence-free survival in patients with stage III melanoma treated with adjuvant immunotherapy | Attrill G, Owen C, Ahmed T, Vergara I, Colebatch A, Conway J, Nahar K, Thompson J, da Silva I, Carlino M, Menzies A, Lo S, Palendira U, Scolyer R, Long G, Wilmott J | 2022 | Journal of Immuno Therapy of Cancer | Immuno-oncology | Cytonuclear, Spatial Analysis, Highplex FL | HALO |
| gp96 Expression in Gliomas and Its Association with Tumor Malignancy and T Cell Infiltrating Level | Li C, Wang Y, Long L, Zhang P, Zhang Y, Ji N | 2022 | Hindawi Journal of Oncology | Immuno-oncology | Spatial Analysis, Highplex FL, Figure Maker | HALO |
| Spartalizumab or placebo in combination with dabrafenib and trametinib in patients with†BRAF†V600-mutant melanoma: exploratory biomarker analyses from a randomized phase 3 trial (COMBI-i) | Tawbi H, Robert C, Brase J, Gusenleitner D, Gasal E, Garrett J, Savchenko A, Gorgun G, Flaherty K, Ribas A, Dummer R, Schadendorf D, Long G, Nathan P, Ascierto P | 2022 | Journal for ImmunoTherapy of Cancer | Oncology | Multiplex IHC, Spatial Analysis | HALO |
| Endothelial cell death after ionizing radiation does not impair vascular structure in mouse tumor models | Kaeppler J, Chen J, Buono M, Vermeer J, Kannan P, Cheng W, Voukantsis D, Thompson J, Hill M, Allen D, Gomes A, Kersemans V, Kinchesh P, Smart S, Buffa F, Nerlov C, Muschel R, Markelc B | 2022 | EMBO Reports | Oncology | Spatial Analysis, Highplex FL | HALO |
| Approaches to spatially resolving the tumour immune microenvironment of hepatocellular carcinoma | Marsh-Wakefield F, Ferguson A, Liu K, Santhakumar C, McCaughan G, Palendira U | 2022 | Therapeutic Advances in Medical Oncology | Review | Multiplex IHC, Spatial Analysis, Highplex FL | HALO |
| Senescence-Associated Molecules and Tumor-Immune-Interactions as Prognostic Biomarkers in Colorectal Cancer | Kellers F, Fernandez A, Konykiewitz B, Schindeldecker M, Tagscherer K, Heintz A, Jesinghaus M, Roth W, Foersch S | 2022 | Frontiers in Medicine | Oncology | Classifier, Multiplex IHC, Spatial Analysis, Registration, TMA | HALO |
| Tau Pathology in Chronic Traumatic Encephalopathy is Primarily Neuronal | Butler M, Dixon E, Stein T, Alvarez V, Huber B, Buckland M, McKee A, Cherry J | 2022 | Journal of Neuropathology & Experimental Neurology | Neuroscience | Multiplex IHC, Spatial Analysis, Highplex FL | HALO, HALO AI |
| Chemoradiation-induced alteration of programmed death-ligand 1, CD8+ tumor-infiltrating lymphocytes and mucin expression in rectal cancer | Baretti M, Zhu Q, Fu W, Meyer J, Wang H, Anders R, Azad N | 2022 | Oncotarget | Oncology | Multiplex IHC, Spatial Analysis | HALO |
| Unveiling the tumor immune microenvironment of organ-specific melanoma metastatic sites | Conway J, Rawson R, Lo S, Ahmed T, Veregara I, Gide T, Attrill G, Carlino M, Saw R, Thompson J, Spillane A, Shannon K, Shivalingam B, Menzies A, Wilmott J, Long G, Scolyer R, da Silva I | 2022 | Journal for ImmunoTherapy of Cancer | Oncology | Classifier, Spatial Analysis, Highplex FL | HALO |
| Haploinsufficiency of the lysosomal sialidase†NEU1†results in a model of pleomorphic rhabdomyosarcoma in mice | Machado E, van de Vlekkert D, Sheppard H, Perry S, Downing S, Laxton J, Ashmun R, Finkelstein D, Neale G, Hu H, Harwood F, Koo S, Grosveld G, d'Azzo A | 2022 | Communications Biology | Oncology | Membrane, Spatial Analysis | HALO |
| Fibrocytes boost tumor-supportive phenotypic switches in the lung cancer niche via the endothelin system | Weigert A, Zheng X, Nenzel A, Turkowski K, Gunther S, Strack E, Sirait-Fischer E, Elwakeel E, Kur I, Nikam V, Valasarajan C, Winter H, Wissgott A, Voswinkel R, Grimminger F, Brune B, Seeger W, Pullamsetti S, Savai R | 2022 | Nature Communications | Oncology | Spatial Analysis, Highplex FL | HALO |
| Use of High-Plex Data Reveals Novel Insights into the Tumour Microenvironment of Clear Cell Renal Cell Carcinoma | De Filippis R, Wolflein G, Um I, Caie P, Warren S, White A, Suen E, To E, Arandjelovic O, Harrison D | 2022 | Cancers | Oncology | Spatial Analysis | HALO, HALO AI |
| Systemic Nos2 Depletion and Cox inhibition limits TNBC disease progression and alters lymphoid cell spatial orientation and density | Somasundaram V, Ridnour L, Cheng R, Walke A, Kedei N, Bhattacharyya D, Wink A, Edmondson E, Butcher D, Warner A, Dorsey T, Scheiblin D, Heinz W, Bryant R, Kinders R, Lipkowitz S, Wong S, Pore M, Hewitt S, McVicar D, Anderson S, Chang J, Glynn S, Ambs S, Lockett S, Wink D | 2022 | Redox Biology | Oncology | Classifer, ISH/FISH, Spatial Analysis, Registration, ISH-IHC/FISH-IF | HALO |
| Annexin A2/TLR2/MYD88 pathway induces arginase 1 expression in tumor-associated neutrophils | Zhang H, Zhu X, Friesen T, Kwak J, Pisarenko T, Mekvanich S, Velasco M, Randolph T, Kargl J, Houghton A | 2022 | Journal of Clinical Investigation | Immuno-oncology | Spatial Analysis, ISH-IHC/FISH-IF | HALO |
| Impact of HPV status on immune responses in head and neck squamous cell carcinoma | Qureshi H, Zhu X, Yang G, Steadele M, Pierce R, Futran N, Lee S, Mendez E, Houghton A | 2022 | Oral Oncology | Oncology | Multiplex IHC, Spatial Analysis, Highplex FL | HALO |
| Developing a Nomogram for Preoperative Prediction of Cervical Cancer Lymph Node Metastasis by Multiplex Immunofluorescence | Wu J, Guo Q, Zhu J, Wu Y, Liang S Chen S, Wang S, Ju X, Wu X | 2022 | Authorea | Oncology | Spatial Analysis, Highplex FL, TMA | HALO |
| Tissue-resident FOLR2+ macrophages associate with CD8+ T cell infiltration in human breast cancer | Ramos R, Missolo-Koussou Y, Gerber-Ferder Y, Bromley C, Bugatti M, Nunez N, Boari J, Richer W, Menger L, Denizeau J, Sedlik C, Caudana P, Kotsias F, Niborski L, Viel S, Bohec M, Lameiras S, Baulande S, Lesage L, Nicolas A, Meseure D, Vincent-Salomon A, Reyal F, Dutertre C, Ginhoux F, Vimeux L, Donnadieu E, Buttard B, Galon J, Zelenay S, Vermi W, Guermonprez P, Piaggio E, Helft J | 2022 | Cell | Immuno-oncology | Classifier, Spatial Analysis, Highplex FL | HALO |
| Neoantigen-specific CD4+ T cells in human melanoma have diverse differentiation states and correlate with CD8+ T cell, macrophage, and B cell function | Veatch J, Lee S, Shasha C, Singhi N, Szeto J, Moshiri A, Kim T, Smythe K, Kong P, Fitzgibbon M, Jesernig B, Bhatia S, Tykodi S, Hall E, Byrd D, Thompson J, Pillarisetty V, Duhen T, Houghton A, Newell E, Gottardo R, Riddell S | 2022 | Cancer Cell | Immuno-oncology | Spatial Analysis, Highplex FL | HALO |
| Escherichia coli-specific CXCL13-producing TFH are associated with clinical efficacy of neoadjuvant PD-1 blockade against muscle-invasive bladder cancer | Goubet A, Lordello L, Silva C, Peguillet I, Gazzano M, Mbogning-Fonkou M, Thelemaque C, Lebacle C, Thibault C, Audenet F, Pignot G, Gravis G, Helissey C, Campbedel L, Roupret M, Xylinas E, Ouzaid I, Dubuisson A, Mazzenga M, Flament C, Ly P, Marty V, Signolle N, Sauvat A, Sbarrato T, Filahi M, Davin C, Haddad G, Khalil J, Bleriot C, Danlos F, Dunsmore G, Mulder K, Silvin A, Raoult T, Archambaud B, Belhechmi S, Boneca I, Cayet N, Moya-Nilges M, Mallet A, Daillere R, Rouleau E, Radulescu C, Allory Y, Fieschi J, Rouanne M, Ginhoux F, Le Teuff G, Derosa L, Marabelle A, Van Dorp J, Van Dijk N, van der Heijden M, Besse B, Andre F, Merad M, Kroemer G, Scoazec J, Zitvogel L, Loriot Y | 2022 | Cancer Discovery | Oncology | Multiplex IHC, Spatial Analysis | HALO |
| Upfront FOLFOXIRI plus bevacizumab with or without atezolizumab in the treatment of patients with metastatic colorectal cancer (AtezoTRIBE): a multicentre, open-label, randomised, controlled, phase 2 trial | Antoniotti C, Rossini D, Pietrantonio F, Catteau A, Salvatore L, Lonardi S, Boquet I, Tamberi S, Marmorino F, Moretto R, Ambrosini M, Tamburini E, Tortora G, Passardi A, Bergamo F, Kassambara A, Sbarrato T, Morano F, Ritorto G, Borelli B, Boccaccino A, Conca V, Giordano M, Ugolini C, Fieschi J, Papadopulos A, Massoue C, Aprile G, Antonyzzo L, Gelsomino F, Martinelli E, Pella N, Masi G, Fontanini G, Boni L, Galon J, Cremolini C | 2022 | The Lancet Oncology | Oncology | Multiplex IHC, Spatial Analysis | HALO |
| Plasma CD27, a surrogate of the intratumoral CD27-CD70 interaction, correlates with immunotherapy resistance in renal cell carcinoma | Benhamouda N, Sam I, Epaillard N, Gey A, Phan L, Pham H, Gruel N, Saldmann A, Pineau J, Hasan M, Quiniou V, Nevoret C, Verkarre V, Libri V, Mella S, Granier C, Broudin C, Ravel P, De Guillebon E, Mauge L, Helley D, Jabla B, Chaput N, Albiges L, Katsahian S, Adam J, Mejean A, Adotevi O, Vano Y, Oudard S, Tartour E | 2022 | Clinical Cancer Research | Oncology | Spatial Analysis, Highplex FL | HALO |
| High endothelial venules associated with better prognosis in esophageal squamous cell carcinoma | Li H, Tang L, Han X, Zhong L, Gao W, Chen Y, Huang J, Wen Z | 2022 | Annals of Diagnostic Pathology | Oncology | Object Colocalization, Spatial Analysis, Highplex FL | HALO |
| Defactinib, pembrolizumab, and gemcitabine in patients with advanced treatment refractory pancreatic cancer: A phase I, dose escalation, and expansion study | Wang-Gillam A, Lim K, McWilliams R, Suresh R, Lockhart A, Brown A, Breden M, Belle J, Herndon J, Bogner S, Pedersen K, Tan B, Boice N, Acharya A, Abdiannia M, Gao F, Yoon H, Zhu M, Trikalinos N, Ratner L, Aranha O, Hawkins W, Herzog B, DeNardo D | 2022 | Clinical Cancer Research | Immuno-oncology | Multiplex IHC, Spatial Analysis | HALO |
| Th1-involved immune infiltrates improve neoadjuvant chemoradiotherapy response of esophageal squamous cell carcinoma | Yuan J, Weng Z, Tan Z, Luo K, Zhong J, Xie X, Qu C, Lin X, Yang H, Wen J, Fu J | 2022 | Cancer Letters | Immuno-oncology | Multiplex IHC, Spatial Analysis | HALO |
| Chronic rhinosinusitis patients display an aberrant immune cell localisation with enhanced S. aureus biofilm metabolic activity and biomass | Shaghayegh G, Cooksley C, Bouras G, Panchatcharam B, Idrizi R, Jana M, Ellis S, Psaltis A, Wormald P, Vreugde S | 2022 | Journal of Allergy and Clinical Immunology | Immunology, Infectious Disease | Classifier, Spatial Analysis, Highplex FL | HALO |
| Autologous T cell therapy for MAGE-A4+ solid cancers in HLA-A*02+ patients: a phase 1 trial | Hong D, Van Tine B, Biswas S, McAlpine C, Johnson M, Olszanski A, Clarke J, Araujo D, Blumenschein G, Kebriaei P, Lin Q, Tipping A, Sanderson J, Wang R, Trivedi T, Annareddy T, Bai J, Rafail S, Sun A, Fernandes L, Navenot J-M, Bushman F, Everett J, Karadeniz D, Broad R, Isabelle M, Naidoo R, Bath N, Betts G, Wolchinsky Z, Batrakou D, Van Winkle E, Elefant E, Ghobadi A, Cashen A, Grand'Maison A, McCarthy P, Fracasso P, Norry E, Williams D, Druta M, Liebner D, Odunsi K, Butler M | 2023 | Nature Medicine | Immuno-oncology | ISH/FISH, Spatial Analysis, Highplex FL | HALO |
| CD8+ Cell Density Gradient across the Tumor Epithelium–Stromal Interface of Non-Muscle Invasive Papillary Urothelial Carcinoma Predicts Recurrence-Free Survival after BCG Immunotherapy | Drachneris J, Rasmusson A, Morkunas M, Fabijonavicius M, Cekauskas A, Jankevicius F, Laurinavicius A | 2023 | Cancers | Immuno-oncology | Multiplex IHC, Spatial Analysis, HALO AI | HALO |
| PD-1+CD8+ T Cells Proximal to PD-L1+CD68+ Macrophages Are Associated with Poor Prognosis in Pancreatic Ductal Adenocarcinoma Patients | Yang X, Wang G, Song Y, Zhuang T, Li Y, Xie Y, Fei X, Zhao Y, Xu D, Hu Y | 2023 | Cancers | Oncology, Immuno-oncology | Classifier, Spatial Analysis, Highplex FL, Registration, TMA | HALO |
| Co-enrichment of CD8-positive T cells and macrophages is associated with clinical benefit of tislelizumab in solid tumors | Ye D, Desai J, Shi J, Liu S, Shen W, Liu T, Shi Y, Wang D, Liang L, Yang S, Ma X, Jin W, Zhang P, Huang R, Shen Z, Zhang Y, Wu Y | 2023 | Biomarker Research | Immuno-oncology | Spatial Analysis, Highplex FL | HALO |
| Cellular mechanisms of heterogeneity in NF2-mutant schwannoma | Chiasson-MacKenzie C, Vitte J, Liu C, Wright E, Flynn E, Stott S, Giovannini M, McClatchey A | 2023 | Nature Communications | Oncology | Classifier, Multiplex IHC, Spatial Analysis, FISH-IF | HALO |
| The localization of molecularly distinct microglia populations to Alzheimer's disease pathologies using QUIVER | Shahidehpour R, Nelson A, Sanders L, Embry C, Nelson P, Bachstetter A | 2023 | Acta Neuropathologica Communications | Neuroscience | Area Quantification, Object Colocalization, Spatial Analysis, Registration | |
| Development of a deep pathomics score for predicting hepatocellular carcinoma recurrence after liver transplantation | Qu W, Tian M, Lu G, Zhou Y, Liu W, Tang Z, Yao Z, Huang R, Zhu G, Jiang X, Tao C, Fang Y, Gao J, Wu X, Chen J, Zhao Q, Yang R, Chu T, Zhou J, Fan J, Yu J, Shi Y | 2023 | Hepatology International | Oncology | Spatial Analysis, Highplex FL | HALO |
| An Immune-Related Gene Expression Signature Predicts Benefit from Adding Atezolizumab to FOLFOXIRI plus Bevacizumab in Metastatic Colorectal Cancer | Antoniotti C, Boccaccino A, Seitz R, Giordano M, Catteau A, Rossini D, Pietrantonio F, Salvatore L, McGregor K, Bergamo F, Conca V, Leonetti S, Morano F, Papiani G, Tamburini E, Bensi M, Murgioni S, Ross D, Passardi A, Boquet I, Nielsen T, Galon J, Varga M, Schweitzer B, and Cremolini C | 2023 | Clinical Cancer Research | Immuno-oncology | Multiplex IHC, Spatial Analysis | HALO |
| Single-cell and spatial transcriptome analysis reveals the cellular heterogeneity of liver metastatic colorectal cancer | Wang F, Long J, Li L, Wu Z, Da T, Wang X, Huang C, Jiang Y, Yao X, Ma H, Lian Z, Zhao Z, Cao J | 2023 | Science Advances | Oncology | Multiplex IHC, Spatial Analysis, Highplex FL | HALO |
| Dissecting tumor lymphocyte infiltration to predict benefit from immune-checkpoint inhibitors in metastatic colorectal cancer: lessons from the AtezoTRIBE study | Moretto R, Rossini D, Catteau A, Antoniotti C, Giordano M, Boccaccino A, Ugolini C, Proietti A, Conca V, Kassambara A, Pietrantonio F, Salvatore L, Lonardi S, Tamberi S, Tamburini E, Poma A, Fieschi J, Fontanini G, Masi G, Galon J, Cremolini C | 2023 | Journal for ImmunoTherapy of Cancer | Immuno-oncology | Multiplex IHC, Spatial Analysis | HALO |
| High tumor mutational burden predicts favorable response to anti-PD-(L)1 therapy in patients with solid tumor: a real-world pan-tumor analysis | Jung J, Heo Y, Park S | 2023 | Journal for ImmunoTherapy of Cancer | Immuno-oncology | Spatial Analysis, Highplex FL | HALO |
| Immunoscore immune checkpoint using spatial quantitative analysis of CD8 and PD-L1 markers is predictive of the efficacy of anti- PD1/PD-L1 immunotherapy in non-small cell lung cancer | Ghiringhelli F, Bibeau F, Greillier L, Fumet J, Ilie A, Monville F, Laugé C, Catteau A, Boquet I, Majdi A, Morgand E, Oulkhouir Y, Brandone N, Adam J, Sbarrato T, Kassambara A, Fieschi J, Garcia S, Lepage A, Tomasini P, Galon J | 2023 | EBioMedicine | Immuno-oncology | Multiplex IHC, Spatial Analysis | HALO |
| Epitope Lability of Phosphorylated Biomarkers of the DNA Damage Response Pathway Results in Increased Vulnerability to Effects of Delayed or Incomplete Formalin Fixation | Wiseman E, Moss J, Atkinson J, Baakza H, Hayes E, Willis S, Waring P, Canales J, Jones G | 2023 | Journal of Histochemistry & Cytochemistry | Other | Classifier, Cytonuclear, Spatial Analysis | HALO |
| The Hippo-YAP signaling pathway drives CD24- mediated immune evasion in esophageal squamous cell carcinoma via macrophage phagocytosis | Yan Z, Hou J, Zhang L, Chen Z, Gao C, Ahmad N, Guo M, Wang W, Han T, Chang T, Kang X, Wang L, Liang Y, Li X, Zhou X | 2023 | Research Square | Immuno-oncology | Spatial Analysis | HALO |
| Stem cell-like T cell depletion in the recurrent head and neck cancer immune microenvironment | Chen L, Lee N, Sarkar R, Katabi N, Li Y, Morris L, Wong R, Sherman E, Reis-Filho J, Hollmann T, Riaz N | 2023 | OncoImmunology | Immuno-oncology | Spatial Analysis, Highplex FL | HALO |
| Differential expression of CD8 defines phenotypically distinct cytotoxic T cells in cancer and multiple sclerosis | Burkard T, Herrero San Juan M, Dreis C, Kiprina A, Namgaladze D, Siebenbrodt K, Luger S, Foerch C, Pfeilschifter J, Weigert A, Radeke H | 2023 | Clinical and Translational Medicine | Oncology, immunology | Classifier, Spatial Analysis, Highplex FL, TMA | HALO |
| Multiplex Immunohistochemistry and Immunofluorescence: A Practical Update for Pathologists | Harms P, Frankel T, Moutafi M, Rao A, Rimm D, Taube J, Thomas D, Chang M, Pantanowitz L | 2023 | Modern Pathology | Review | Spatial Analysis, Highplex FL | HALO |
| Neighboring macrophage-induced alteration in the phenotype of colorectal cancer cells in the tumor budding area | Kawamura I, Ohe R, Suzuki K, Kabasawa T, Kitoaka T, Takahara D, Kono M, Uchiyama N, Musha H, Futakuchi M, Motoi F | 2023 | Research Square | Immuno-oncology | Multiplex IHC, Spatial Analysis | HALO |
| Single-cell RNA-sequencing reveals heterogeneity and intercellular crosstalk in human tuberculosis lung | Wang, H, Wen, Z, Niu, L, Chen, X, Liu, H, Zhang, S, Xu, J, Zhu, Y, Li, H, Chen, H, Shi, L, Wan, L, Li, L, Li, M, Wong, K, Song, Y | 2023 | Journal of Infection | Immunology, Infectious Disease | Spatial Analysis, Highplex FL | HALO |
| Tumor-associated macrophages trigger MAIT cell dysfunction at the HCC invasive margin | Ruf B, Bruhns M, Babaei S, Kedei N, Ma L, Revsine M, Benmebarek MR, Ma C, Heinrich B, Subramanyam V, Qi J, Wabitsch S, Green BL, Bauer KC, Myojin Y, Greten LT, McCallen JD, Huang P, Trehan R, Wang X, Nur A, Soika DQM, Pouzolles M, Evans CN, Chari R, Kleiner DE, Telford W, Dadkhah K, Ruchinskas A, Stovroff MK, Kang J, Oza K, Ruchirawat M, Kroemer A, Wang XW, Claassen M, Korangy F, Greten TF | 2023 | Cell | Immuno-oncology | Classifier, Spatial Analysis, Highplex FL, Figure Maker | HALO |
| Tumor cell HLA class I expression and pathologic response following neoadjuvant immunotherapy for newly diagnosed head and neck cancer | Robbins Y, Friedman J, Redman J, Sievers C, Lassoued W, Gulley J, Allen C | 2023 | Oral Oncology | Immuno-oncology | Classifier, Spatial Analysis, Highplex FL | HALO |
| Anticoagulants Enhance Molecular and Cellular Immunotherapy of Cancer by Improving Tumor Microcirculation Structure and Function and Redistributing Tumor Infiltrates | Wei F, Su Y, Quan Y, Li X, Zou Q, Zhang L, Li S, Jiang M, Lin G, Liang P, He J, Xie K | 2023 | Clinical Cancer Research | Immuno-oncology | Classifier, Multiplex IHC, Spatial Analysis, Highplex FL | HALO |
| Spatial mapping reveals granuloma diversity and histopathological superstructure in human tuberculosis | Sawyer A, Patrick E, Edwards J, Wilmott J, Fielder T, Yang Q, Barber D, Ernst J, Britton W, Palendira U, Chen X, Feng C | 2023 | Journal of Experimental Medicine | Immunology | Multiplex IHC, Spatial Analysis, Highplex FL | HALO |
| Spatial Proximity and Relative Distribution of Tumor-Infiltrating Lymphocytes and Macrophages Predict Survival in Melanoma | De Logu F, Ugolini F, Iannone L F, Simi S, Maio V, de Giorgi V, di Giacomo A M, Miracco C, Cossu A, Palmieri G, Mandalà M, Massi D | 2023 | Laboratory Investigation | Oncology | Multiplex IHC, Spatial Analysis | HALO, HALO Link |
| Histological spatial analysis on the induction of PD-L1+ macrophages by CD8+ T cells at the marginal microenvironment of triple-negative breast cancer | Suzuki K, Ohe R, Kabasawa T, Kitaoka T, Kawai M, Motoi F, Futakuchi M | 2023 | Breast Cancer | Oncology | Multiplex IHC, Spatial Analysis | HALO |
| A neutrophil extracellular traps-related classification predicts prognosis and response to immunotherapy in colon cancer | Feng C, Li Y, Tai Y, Zhang W, Wang H, Lian S, Jin-si-han E, Liu Y, Li X, Chen Q, He M, Lu Z | 2023 | Scientific Reports | Immuno-oncology | Spatial Analysis, Highplex FL, Figure Maker, TMA | HALO |
| Specific Polo-Like Kinase 1 Expression in Nodular Lymphocyte-Predominant Hodgkin Lymphoma Suggests an Intact Immune Surveillance Program | Weiss J, Gibbons K, Ehyaee V, Perez-Silos V, Zevallos A, Maienschein-Cline M, Brister E, Sverdlov M, Shah E, Balakrishna J, Symes E, Frederiksen J K, Gann P H, Post R, Lopez-Hisijos N, Reneau J, Venkataraman G, Bailey N, Brown N A, Xu M L, Murga-Zamalloa C | 2023 | The American Journal of Pathology | Oncology | Area Quantification, Spatial Analysis | HALO |
| The tumor-derived cytokine Chi3l1 induces neutrophil extracellular traps that promote T cell exclusion in triple-negative breast cancer | Taifour T, Attalla SS, Zuo D, Gu Y, Sanguin-Gendreau V, Proud H, Solymoss E, Bui T, Kuasne H, Papavasiliou V, Lee CG, Kamle S, Siegel PM, Elias JA, Park M, Muller WJ | 2023 | Immunity | Immuno-oncology | Classifier, Multiplex IHC, Spatial Analysis, Highplex FL | HALO |
| Components of the tumor immune microenvironment based on m-IHC correlate with prognosis and subtype of triple-negative breast cancer | Lin L, Li H, Wang X, Wang Z, Su G, Zhou J, Sun S, Ma X, Chen Y, You C, Gu Y | 2023 | Cancer Medicine | Immuno-oncology | Classifier, Spatial Analysis, Highplex FL | HALO |
| Analysis of off-tumour toxicities of T-cell-engaging bispecific antibodies via donor-matched intestinal organoids and tumouroids | Harter MF, Recaldin T, Gerard R, Avignon B, Bollen Y, Esposito C, Guja-Jarosz K, Kromer K, Filip A, Aubert J, Schneider A, Bacac M, Bscheider M, Stokar-Regenscheit N, Piscuoglio S, Beumer J, Gjorevski N | 2023 | Nature Biomedical Engineering | Immuno-oncology | Spatial Analysis, Highplex FL | HALO, HALO AI |
| Integrated multi-omics profiling to dissect the spatiotemporal evolution of metastatic hepatocellular carcinoma | Sun Y, Wu P, Zhang Z, Wang Z, Zhou K, Song M, Ji Y, Zang F, Lou L, Rao K, Wang P, Gu Y, Gu J, Lu B, Chen L, Pan X, Zhao X, Peng L, Liu D, Chen X, Wu K, Lin P, Wu L, Su Y, Du M, Hou Y, Yang X, Qiu S, Shi Y, Sun H, Zhou J, Huang X, Peng D, Zhang L, Fan J | 2023 | Cancer Cell | Oncology | Classifier, Multiplex IHC, Spatial Analysis, Highplex FL | HALO |
| Targeting myeloid chemotaxis to reverse prostate cancer therapy resistance | Guo C, Sharp A, Gurel B, Crespo M, Figueiredo I, Jain S, Vogl U, Rekowski J, Rouhifard M, Gallagher L, Yuan W, Carreira S, Chandran K, Paschalis A, Colombo I, Stathis A, Bertan C, Seed G, Goodall J, Raynaud F, Ruddle R, Swales K, Malia J, Bogdan D, Tiu C, Caldwell R, Aversa C, Ferreira A, Neeb A, Tunariu N, Westaby D, Carmichael J, Maza M, Yap C, Matthews R, Badham H, Prout T, Turner A, Parmar M, Tovey H, Riisnaes R, Flohr P, Gil J, Waugh D, Decordova S, Schlag A, Calì B, Alimonti A, de Bono J S | 2023 | Nature | Oncology | Spatial Analysis, Highplex FL | HALO, HALO AI |
| Gastric tubular adenocarcinoma with diffuse neutrophils infiltrating: characteristics and probable treatment strategy | Wang B, Zhu Y, Wang S, Li Z, Wang L, Rao W, Cheng N, Chen R, Ying J, Xue L | 2023 | Gastric Cancer | Oncology | Multiplex IHC, Spatial Analysis, Registration | HALO |
| Neoadjuvant chemotherapy is linked to an amended anti-tumorigenic microenvironment in gastric cancer | Huan X, Zou K, Zhang P, Ding H, Luo C, Xiang C, Xu S, Zhuang Y, Wu C, Wang Y, Wu X, Chen C, Zhang J, Yao X, Liu F, Liu S, Wu Z | 2023 | International Immunopharmacology | Immuno-oncology | Spatial Analysis, Highplex FL | HALO |
| Analysis of off-tumour toxicities of T-cell-engaging bispecific antibodies via donor-matched intestinal organoids and tumouroids | Harter MF, Recaldin T, Gerard R, Avignon B, Bollen Y, Esposito C, Guja-Jarosz K, Kromer K, Filip A, Aubert J, Schneider A, Bacac M, Bscheider M, Stokar-Regenscheit N, Piscuoglio S, Beumer J, Gjorevski N | 2023 | Nature Biomedical Engineering | Immuno-oncology | Spatial Analysis, Highplex FL | HALO, HALO AI |
| Integrated multi-omics profiling to dissect the spatiotemporal evolution of metastatic hepatocellular carcinoma | Sun Y, Wu P, Zhang Z, Wang Z, Zhou K, Song M, Ji Y, Zang F, Lou L, Rao K, Wang P, Gu Y, Gu J, Lu B, Chen L, Pan X, Zhao X, Peng L, Liu D, Chen X, Wu K, Lin P, Wu L, Su Y, Du M, Hou Y, Yang X, Qiu S, Shi Y, Sun H, Zhou J, Huang X, Peng D, Zhang L, Fan J | 2023 | Cancer Cell | Oncology | Classifier, Multiplex IHC, Spatial Analysis, Highplex FL | HALO |
| Targeting myeloid chemotaxis to reverse prostate cancer therapy resistance | Guo C, Sharp A, Gurel B, Crespo M, Figueiredo I, Jain S, Vogl U, Rekowski J, Rouhifard M, Gallagher L, Yuan W, Carreira S, Chandran K, Paschalis A, Colombo I, Stathis A, Bertan C, Seed G, Goodall J, Raynaud F, Ruddle R, Swales K, Malia J, Bogdan D, Tiu C, Caldwell R, Aversa C, Ferreira A, Neeb A, Tunariu N, Westaby D, Carmichael J, Maza M, Yap C, Matthews R, Badham H, Prout T, Turner A, Parmar M, Tovey H, Riisnaes R, Flohr P, Gil J, Waugh D, Decordova S, Schlag A, Calì B, Alimonti A, de Bono J S | 2023 | Nature | Oncology | Spatial Analysis, Highplex FL | HALO, HALO AI |
| Gastric tubular adenocarcinoma with diffuse neutrophils infiltrating: characteristics and probable treatment strategy | Wang B, Zhu Y, Wang S, Li Z, Wang L, Rao W, Cheng N, Chen R, Ying J, Xue L | 2023 | Gastric Cancer | Oncology | Multiplex IHC, Spatial Analysis, Registration | HALO |
| Neoadjuvant chemotherapy is linked to an amended anti-tumorigenic microenvironment in gastric cancer | Huan X, Zou K, Zhang P, Ding H, Luo C, Xiang C, Xu S, Zhuang Y, Wu C, Wang Y, Wu X, Chen C, Zhang J, Yao X, Liu F, Liu S, Wu Z | 2023 | International Immunopharmacology | Immuno-oncology | Spatial Analysis, Highplex FL | HALO |
| Neoadjuvant radioimmunotherapy in pancreatic cancer enhances effector T cell infiltration and shortens their distances to tumor cells | WANG J, GAI J, ZHANG T, NIU N, QI H, THOMAS II D L, LI K, XIA T, RODRIGUEZ C, PARKINSON R, DURHAM J, MCPHAUL T, NARANG A K, ANDERS R A, OSIPOV A, WANG H, HE J, LAHERU D A, HERMAN J M, LEE V, JAFFEE E M, THOMPSON E D, ZHU Q, ZHENG L | 2024 | Science Advances | Immuno-oncology | Spatial Analysis, Highplex FL, Registration | HALO |
| Single-cell and spatial multi-omics highlight effects of anti-integrin therapy across cellular compartments in ulcerative colitis | Mennillo E, Kim Y, Lee G, Rusu I, Patel R K, Dorman L C, Flynn E, Li S, Bain J L, Andersen C, Rao A, Tamaki S, Tsui J, Shen A, Lotstein M L, Rahim M, Naser M, Bernard-Vazquez F, Eckalbar W, Cho S, Beck K, El-Nachef N, Lewin S, Selvig D R, Terdiman J P, Mahadevan U, Oh D Y, Fragiadakis G K, Pisco A, Combes A J, Kattah M G | 2024 | Nature Communications | Immunology | ISH, Spatial Analysis, Highplex FL, TMA | HALO, HALO AI |
| Serum amyloid A promotes glycolysis of neutrophils during PD-1 blockade resistance in hepatocellular carcinoma | He M, Liu Y, Chen S, Deng H, Feng C, Qiao S, Chen Q, Hu Y, Chen H, Wang X, Jiang X, Xia X, Zhao M, Lyu N | 2024 | Nature Communications | Immuno-oncology | Spatial Analysis, Highplex FL | HALO |
| PDGFRa+ ITGA11+ fibroblasts foster early-stage cancer lymphovascular invasion and lymphatic metastasis via ITGA11-SELE interplay | Zheng H, An M, Luo Y, Diao X, Zhong W, Pang M, Lin Y, Chen J, Li Y, Kong Y, Zhao Y, Yin Y, Ai L, Huang J, Chen C, Lin T | 2024 | Cancer Cell | Oncology | Spatial Analysis, Highplex FL | HALO |
| Multi-omics and imaging mass cytometry characterization of human kidneys to identify pathways and phenotypes associated with impaired kidney function. | Asowata E O, Romoli S, Sargeant R, Tan J Y, Hoffmann S, Huang M M, Mahbubani K T, Krause F N, Jachimowicz D, Agren R, Koulman A, Jenkins B, Musial B, Griffin J L, Soderberg M, Ling S, Hansen P B L, Saeb-Parsy K, Woollard K J | 2024 | Kidney International | Other | ISH, Spatial Analysis, Highplex FL | HALO, HALO AI |
Related HALO Modules
Quantify expression of up to five brightfield stains in any cellular compartment - membrane, nucleus or cytoplasm.
Learn MoreQuantify expression of an unlimited number of biomarkers in any cellular compartment - membrane, nucleus or cytoplasm.
Learn MoreSeparate multiple tissue classes across a tissue using a learn-by-example approach. Can be used in conjunction with all other modules (fluorescent and brightfield) to select specific tissue classes for further analysis.
Learn MoreUse the arrows above to view additional related modules
Want to Learn More?
Fill out the form below to request information about any of our software products.
You can also drop us an email at info@indicalab.com