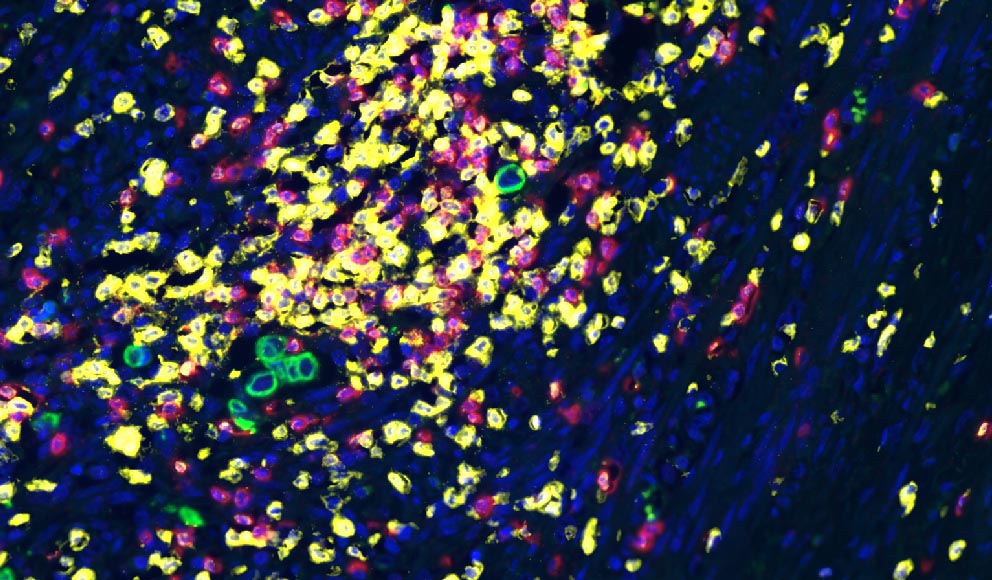
Automated Analysis of Lymphocytic Infiltration, Tumor Budding, and Their Spatial Relationship Improves Prognostic Accuracy in Colorectal Cancer
Both immune profiling and tumor budding significantly correlate with colorectal cancer patient outcome but are traditionally reported independently. This study evaluated the association and interaction between lymphocytic infiltration and tumor budding, coregistered on a single slide, in order to determine a more precise prognostic algorithm for patients with stage II colorectal cancer. Multiplexed immunofluorescence and automated image analysis were used for the quantification of CD3þCD8þ T cells, and tumor buds (TBs), across whole slide images of three independent cohorts (training cohort: n¼114, validation cohort 1: n¼56, validation cohort 2: n ¼ 62). Machine learning algorithms were used for feature selection and prognostic risk model development. High numbers of TBs [HR ¼ 5.899; 95% confidence interval (CI) 1.875– 18.55], low CD3þ T-cell density (HR ¼ 9.964; 95% CI, 3.156–31.46), and low mean number of CD3þCD8þ T cells within 50 mm of TBs (HR ¼ 8.907; 95% CI, 2.834–28.0) were associated with reduced disease-specific survival. A prognostic signature, derived from integrating TBs, lymphocyte infiltration, and their spatial relationship, reported a more significant cohort stratification (HR ¼ 18.75; 95% CI, 6.46–54.43), than TBs, Immunoscore, or pT stage. This was confirmed in two independent validation cohorts (HR ¼ 12.27; 95% CI, 3.524–42.73; HR ¼ 15.61; 95% CI, 4.692–51.91). The investigation of the spatial relationship between lymphocytes and TBs
within the tumor microenvironment improves accuracy of prognosis of patients with stage II colorectal cancer through an automated image analysis and machine learning workflow.
Ines P. Nearchou, Kate Lillard, Christos G. Gavriel, Hideki Ueno, David J. Harrison, and Peter D. Caie
Cancer Immunology Research| Published March 7, 2019 | DOI 10.1158/2326-6066.CIR-18-0377